 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010921494.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 318aa MW: 34996.1 Da PI: 4.8894 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 30.6 | 5.9e-10 | 253 | 285 | 23 | 55 |
SS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 23 rypsaeereeLAkklgLterqVkvWFqNrRake 55
+yp++ee+++LA+ +gL+++q+ +WF N+R ++
XP_010921494.1 253 PYPTEEEKAKLAEMTGLDQKQINNWFINQRKRH 285
8*****************************885 PP
| |||||||
| 2 | ELK | 35.1 | 2.9e-12 | 205 | 226 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELK++Ll+KYsgyL++L++EF+
XP_010921494.1 205 ELKEMLLKKYSGYLSNLRKEFL 226
9********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 3.5E-21 | 58 | 102 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 1.2E-20 | 60 | 100 | IPR005540 | KNOX1 |
| SMART | SM01256 | 7.4E-26 | 105 | 156 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 1.2E-24 | 108 | 155 | IPR005541 | KNOX2 |
| PROSITE profile | PS51213 | 11.261 | 205 | 225 | IPR005539 | ELK domain |
| Pfam | PF03789 | 1.1E-9 | 205 | 226 | IPR005539 | ELK domain |
| SMART | SM01188 | 1.9E-6 | 205 | 226 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 13.038 | 225 | 288 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 3.25E-20 | 227 | 297 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 1.5E-14 | 227 | 292 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 4.9E-27 | 230 | 290 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.32E-13 | 237 | 289 | No hit | No description |
| Pfam | PF05920 | 1.2E-17 | 245 | 284 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 263 | 286 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010073 | Biological Process | meristem maintenance | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 318 aa Download sequence Send to blast |
MEDLYSIHPV ISRGGEVAAV GSASSFAYLE ADRSQGGETA EASDISAGGG GGGSDLTDLI 60 KAQIANHPRY PSLVAAYIEC RKVGAPPEMA SLLEEIGRER YTSAGCGEIG ADPELDEFME 120 SYCRVLQRYK EELSKPFNEA ASFLNSIEMQ LSNLCKGRTT SSSTTAATGN SPSDEVVGSS 180 EEELSCGDVD ASECQESGSR LADHELKEML LKKYSGYLSN LRKEFLKKRK KGKLPKDARL 240 TLLDWWHTHY RWPYPTEEEK AKLAEMTGLD QKQINNWFIN QRKRHWKPSE DMRFALMEGV 300 SGGSSGTMLY FDTGTVGP |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 220 | 229 | LRKEFLKKRK |
| 2 | 226 | 230 | KKRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may be involved in shoot formation during early embryogenesis. {ECO:0000269|PubMed:10488233}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00670 | PBM | Transfer from GRMZM2G087741 | Download |
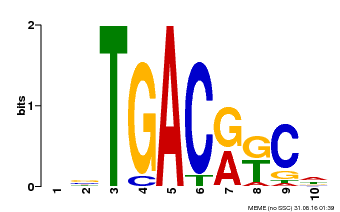
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010921494.1 | 0.0 | homeobox protein knotted-1-like 1 | ||||
| Swissprot | Q9FP29 | 1e-131 | KNOS1_ORYSJ; Homeobox protein knotted-1-like 1 | ||||
| TrEMBL | F2YNF2 | 0.0 | F2YNF2_ELAGV; Knotted-like homebox protein | ||||
| STRING | XP_008781110.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1698 | 38 | 92 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G23380.1 | 2e-82 | KNOTTED1-like homeobox gene 6 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



