 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010916479.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 349aa MW: 38341.5 Da PI: 9.8197 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 38.9 | 2.1e-12 | 55 | 103 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s+ykGV + +grW A+I++ ++r++lg+f + +Aa+a++ a+++++g
XP_010916479.1 55 SQYKGVVPQP-NGRWGAQIYEK-----HQRVWLGTFADEADAARAYDVAAQRFRG 103
78****9888.8*********3.....5**********99*************98 PP
| |||||||
| 2 | B3 | 95.6 | 3.4e-30 | 179 | 278 | 1 | 96 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrse 89
f+kv tpsdv+k++rlv+pk++ae++ + ++ l +ed g++W+++++y+++s++yvltkGW++Fvk++ Lk+gD+v F +++ e
XP_010916479.1 179 FDKVVTPSDVGKLNRLVIPKQHAEKNfplkigS-G-SAGMLLNFEDVIGKVWRFRYSYWNSSQSYVLTKGWSRFVKEKSLKAGDVVSFWRSRGPE 271
89***************************9743.3.3689************************************************9887777 PP
E..EEEE CS
B3 90 felvvkv 96
++l++ +
XP_010916479.1 272 KQLYIDW 278
7777766 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 5.15E-23 | 55 | 110 | No hit | No description |
| SuperFamily | SSF54171 | 9.81E-17 | 55 | 111 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 2.7E-20 | 56 | 111 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 5.0E-7 | 56 | 103 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.757 | 56 | 111 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 5.1E-29 | 56 | 117 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 1.6E-40 | 170 | 285 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 3.27E-31 | 175 | 279 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.09E-28 | 178 | 272 | No hit | No description |
| PROSITE profile | PS50863 | 13.578 | 179 | 280 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 5.3E-27 | 179 | 278 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 2.1E-26 | 179 | 282 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 349 aa Download sequence Send to blast |
MDGSCVEELS NESGKRPPPA AAAPLYRMGS GACSVVLDPT TESGVEAESR KLPSSQYKGV 60 VPQPNGRWGA QIYEKHQRVW LGTFADEADA ARAYDVAAQR FRGRDAVTNF TPLSENDDDG 120 AAELGFLAAH SKAEIVDMLR KHTYCDELRQ SKRAGNASGS IVGKRSPPSS SVSDREILFD 180 KVVTPSDVGK LNRLVIPKQH AEKNFPLKIG SGSAGMLLNF EDVIGKVWRF RYSYWNSSQS 240 YVLTKGWSRF VKEKSLKAGD VVSFWRSRGP EKQLYIDWKP RAAAAAAFAV RPVGHVQVVR 300 LFGVNIFRTP VGCGAAAGNG CGDREGKRVR EMELFPPEQF VKKQCIRAL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 6e-52 | 176 | 284 | 11 | 120 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). Transcriptional repressor of flowering time on long day plants. Acts directly on FT expression by binding 5'-CAACA-3' and 5'-CACCTG-3 sequences (Probable). Functionally redundant with TEM1. {ECO:0000250, ECO:0000269|PubMed:18718758, ECO:0000305}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
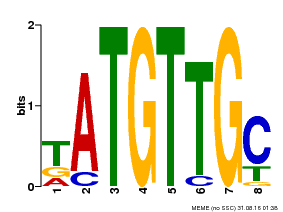
| Motif ID | Method | Source | Motif file |
| MP00025 | PBM | Transfer from AT1G68840 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010916479.1 | 0.0 | AP2/ERF and B3 domain-containing transcription repressor RAV2 | ||||
| Swissprot | P82280 | 1e-135 | RAV2_ARATH; AP2/ERF and B3 domain-containing transcription repressor RAV2 | ||||
| TrEMBL | A0A2H3YIN7 | 0.0 | A0A2H3YIN7_PHODC; AP2/ERF and B3 domain-containing transcription repressor RAV2-like | ||||
| STRING | XP_008799316.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP456 | 38 | 203 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G68840.2 | 1e-121 | related to ABI3/VP1 2 | ||||



