 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010915409.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 838aa MW: 92063.5 Da PI: 6.4631 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59 | 7.5e-19 | 18 | 76 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++ +++ps +r++L +++ +++ +q+kvWFqNrR +ek+
XP_010915409.1 18 KYVRYTPEQVEALERLYHDCPKPSSMRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 76
5679*****************************************************97 PP
| |||||||
| 2 | START | 182.5 | 2.4e-57 | 162 | 370 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEEEEE CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..galqlm 94
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v m+++ v+e+l+d++ W ++++++++++v+ g g+++l
XP_010915409.1 162 IAEETLTEFLSKATGTAVEWVQMPGMKPGPDSIGIVAISHGCTGVAARACGLVGMEPT-KVAEILKDRPSWFRDCRSVDVINVLPAGnsGTIELL 255
789*******************************************************.8888888888************************** PP
EEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHH CS
START 95 vaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslv 185
+++l+a+++l+p Rdf+ +Ry+ l++g++v++++S++s+q p+ +++vRae+lpSg+li+p+++g+s +++v+h+dl+ ++++++lr+l+
XP_010915409.1 256 YMQLYAPTTLAPaRDFWLLRYTSALEDGSLVVCERSLNSTQGGPSmpaVQPFVRAEMLPSGYLIRPCEGGGSIIHIVDHMDLEPWSIPEVLRPLY 350
********************************************999899********************************************* PP
HHHHHHHHHHHHHHTXXXXX CS
START 186 ksglaegaktwvatlqrqce 205
+s+++ ++kt++a+l+++++
XP_010915409.1 351 ESSTVLAQKTTMAALRHLRQ 370
***************99876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.564 | 13 | 77 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.1E-16 | 15 | 81 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 5.69E-17 | 16 | 78 | No hit | No description |
| SuperFamily | SSF46689 | 7.27E-17 | 17 | 81 | IPR009057 | Homeodomain-like |
| Pfam | PF00046 | 1.8E-16 | 18 | 76 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 2.1E-18 | 20 | 76 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 1.14E-6 | 70 | 109 | No hit | No description |
| PROSITE profile | PS50848 | 26.319 | 152 | 366 | IPR002913 | START domain |
| CDD | cd08875 | 1.29E-82 | 156 | 371 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 5.5E-25 | 159 | 367 | IPR023393 | START-like domain |
| SMART | SM00234 | 3.5E-42 | 161 | 371 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 7.42E-39 | 161 | 372 | No hit | No description |
| Pfam | PF01852 | 3.3E-55 | 162 | 370 | IPR002913 | START domain |
| Pfam | PF08670 | 1.9E-51 | 695 | 837 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 838 aa Download sequence Send to blast |
MMAVSSGKEG KTAMDPGKYV RYTPEQVEAL ERLYHDCPKP SSMRRQQLIR ECPILSNIEP 60 KQIKVWFQNR RCREKQRKEA SRLQAVNRKL TAMNKLLMEE NDRLQKQVSQ LVYENGYFRQ 120 QTRNAALTTP DTSCESVVTS GQHHLTAQRP PRDASPAGLM SIAEETLTEF LSKATGTAVE 180 WVQMPGMKPG PDSIGIVAIS HGCTGVAARA CGLVGMEPTK VAEILKDRPS WFRDCRSVDV 240 INVLPAGNSG TIELLYMQLY APTTLAPARD FWLLRYTSAL EDGSLVVCER SLNSTQGGPS 300 MPAVQPFVRA EMLPSGYLIR PCEGGGSIIH IVDHMDLEPW SIPEVLRPLY ESSTVLAQKT 360 TMAALRHLRQ IVHEVSQSTV TGWGRRPAAL RALSQRLSRG FNEALNGFAD DGWSMISGDG 420 TDDVTILVNS SPSRLMGLNL GFTNGFPSLS NAVLCAKASM LLQNVSPAML LRFVREHRSE 480 WADSNIDAYS AAAVRASPCT LPMSRIGCFG GQVILPLAHT LEHEEFLEVL KLENTGRSQD 540 ALMARDLFLL QLCSGVDENA VGTCSELVFA PIDASFADDA PLLPSGFRII PLDSVMENSS 600 PNRTLDLASA LEIGSTGNRV SGDYSGNSTS MRSVMTIAFQ FAFESHLQEN VACMARQYIR 660 NIISSVQRVA LALSPSHLGS HGGLRPPPGT PEAVTLVRWI CYSYRGHSGV ELLKTTTEGS 720 ESLLKILWHH SDAIICCSLK AMPVFTFANQ AGLDMLETTL VALQDITLEK IFDDQGRKTL 780 CTEFPHIMQQ GFACLQGGLC ISSMGRPVSY DSAVAWKVLN DEDEAHCICF MFVNWSFV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
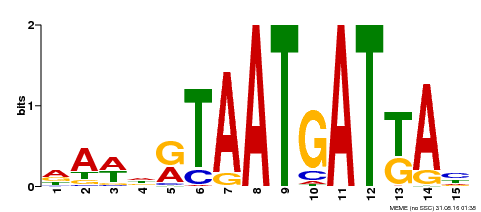
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010915409.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 isoform X1 | ||||
| Refseq | XP_010915410.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 isoform X1 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A2H3Y0I2 | 0.0 | A0A2H3Y0I2_PHODC; homeobox-leucine zipper protein ATHB-15-like | ||||
| STRING | XP_008791633.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP374 | 38 | 197 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



