 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010912370.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 974aa MW: 107291 Da PI: 6.9886 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 92.4 | 3.2e-29 | 608 | 659 | 1 | 52 |
RWP-RK 1 aekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
aek+isle+l++yFs+++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rki+++
XP_010912370.1 608 AEKSISLEVLQQYFSGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKISKV 659
589**********************************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.7 | 598 | 679 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 3.2E-26 | 612 | 659 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 1.6E-22 | 872 | 965 | No hit | No description |
| SMART | SM00666 | 1.8E-21 | 879 | 961 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 5.6E-24 | 879 | 961 | No hit | No description |
| PROSITE profile | PS51745 | 16.06 | 879 | 961 | IPR000270 | PB1 domain |
| CDD | cd06407 | 2.56E-33 | 880 | 960 | No hit | No description |
| Pfam | PF00564 | 8.2E-14 | 880 | 960 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 974 aa Download sequence Send to blast |
MESPPFIPVA SCPPLXSPSS HVGNRRREAA RRRLDSPRPG AHGPRSRHRR LHLALRSQRL 60 RLHSSLLFPS LLLLLPSPWP PSAFLPPSPS PIWIVEDRAA EISSFLEDCP RFLNANEGAR 120 CGNVNASNSK ARRLPLPAVE ENADDSRVIK ERMTQALRHF KESTDQHVLV QVWAPMKKGD 180 RCFLTTSGQP FVLDPQSTRL LQYRTVSVMY IFSVDEDGEG SLGLPGRVYR HRLPEWTPNV 240 QYYSSKEYPR LAHALNYNVQ GSLALPVFEP SGQSCVGVVE LVMTLPKINY AREVDKVCKA 300 LQAVNLKSSE ILDHPNIQIS NEGRQEALAE ILEVLTVVCE AQKLPLAQTW VPCKRRTILA 360 HGGGLKKICS SFDGSCMGQV CMSTTDVAFY VIDAHMWGFR EACVEHHLQI GQGVAGRAFA 420 LRKPCFAEDV TKFSKSEYPL VHYARMFGLV GCLAICLQST HSGDDDYILE FFLPHDCKYL 480 IEQQALLDSI SALMKQCFRS LKVVSDLELQ KGISFEIINM FANENHGFEP KHICNQICDS 540 HLYASPETYT LEGYDDSQNE GNVTSDSRVQ HFPTHANAEK ASNIHFDSAG SLLSNRNNGA 600 PEKRRGKAEK SISLEVLQQY FSGSLKDAAK SLGVCPTTMK RICRQHGISR WPSRKISKVN 660 RSLSKLKRVI ESVQVAEGAF HLTSLGCPLP VAVGSISWPV NLDGSKEITG AVKDNVSSPH 720 KTPGKNDQHD KLLAQQELGH DDSSPMPEPG KDSPCSKTNG SSEGSMDTPA SLGSFHESPV 780 NETFLDDASP PSDLKQGIVG DSLEFTLQHA GGPRLPPVCL MPGVNVAKPQ PQLAGMLIED 840 SGSSKDLKNL CSNARDGCQD EHVMAANPLN PMAMPETRMV TIKASHQEDI IRFRLACTAG 900 IVALRDEVTK RLKLEVGTFD IKYLDDDHEW VMLACNADLE ECIEISGLSG SHVIRLAVHD 960 VTTHLGSSCA SLGD |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
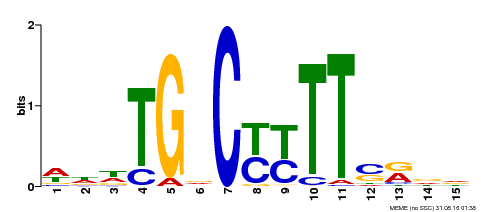
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010912370.2 | 0.0 | LOW QUALITY PROTEIN: protein NLP3 | ||||
| Swissprot | Q5NB82 | 0.0 | NLP3_ORYSJ; Protein NLP3 | ||||
| TrEMBL | A0A2H3X1V9 | 0.0 | A0A2H3X1V9_PHODC; protein NLP3-like isoform X1 | ||||
| STRING | XP_008777047.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4778 | 36 | 60 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||



