 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010906065.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 317aa MW: 34838.8 Da PI: 5.8612 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 45.9 | 1.3e-14 | 26 | 73 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g W++eEd++l+ + G g+W+ +ar g+ R++k+c++rw +yl
XP_010906065.1 26 KGLWSPEEDDKLISYMLSNGQGCWSDVARNAGLQRCGKSCRLRWINYL 73
678*******************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 49.5 | 9.7e-16 | 79 | 122 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
rg+++++E+e +++++ lG++ W+ Ia++++ gRt++++k++w++
XP_010906065.1 79 RGAFSPQEEEVIIHLHSILGNR-WSQIAARLP-GRTDNEIKNFWNS 122
89********************.*********.***********97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 15.665 | 21 | 73 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.09E-27 | 24 | 120 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.6E-11 | 25 | 75 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.9E-13 | 26 | 73 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.9E-22 | 27 | 80 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 5.72E-9 | 29 | 73 | No hit | No description |
| PROSITE profile | PS51294 | 25.202 | 74 | 128 | IPR017930 | Myb domain |
| SMART | SM00717 | 5.9E-15 | 78 | 126 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.0E-14 | 79 | 122 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.87E-10 | 81 | 121 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.0E-25 | 81 | 129 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:2000652 | Biological Process | regulation of secondary cell wall biogenesis | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 317 aa Download sequence Send to blast |
MRKPDPPAKN NNMSNNNSNG ATKLKKGLWS PEEDDKLISY MLSNGQGCWS DVARNAGLQR 60 CGKSCRLRWI NYLRPDLKRG AFSPQEEEVI IHLHSILGNR WSQIAARLPG RTDNEIKNFW 120 NSTIKKRLKN SSSPSPNHSG SPEPKVGMGG LTTLNDHEMK TMCMDSASSS SSSTMQGLSI 180 SNQYDLLPLP EVNCGMTGLG TSYFHEPTSF PQVGFGNGNY GSHGMMMGGG LMGGGCELFV 240 PALESTSTEE NGTSQNTTRN TSDPCHSHNN DVNKINSDAA VGNEIYWEGA NIRMEEWDLD 300 DLMKDVSSFP FLDFQAE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 5e-28 | 20 | 128 | 1 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that acts as molecular switch in the NAC012/SND1-mediated transcriptional network regulating secondary wall biosynthesis. Is directly activated by NAC012/SND1 and its close homologs, including NAC043/NST1, NAC066/NST2, NAC101/VND6 and NAC030/VND7. Is required for functional expression of a number of secondary wall-associated transcription factors and secondary wall biosynthetic genes involved in cellulose, xylan and lignin synthesis. Functions redundantly with MYB46 in the transcriptional regulatory cascade leading to secondary wall formation in fibers and vessels (PubMed:19808805). Transcription activator that binds to the DNA consensus sequence 5'-ACC[AT]A[AC][TC]-3', designated as the secondary wall MYB-responsive element (SMRE). Regulates directly numerous transcription factors and a number of genes involved in secondary wall biosynthesis that contain SMRE elements in their promoters (PubMed:22197883). {ECO:0000269|PubMed:19808805, ECO:0000269|PubMed:22197883}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00336 | DAP | Transfer from AT3G08500 | Download |
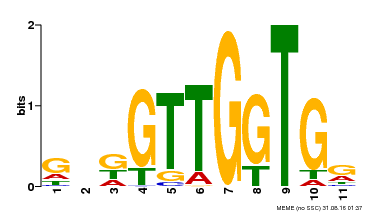
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010906065.1 | 0.0 | transcription factor MYB83 | ||||
| Swissprot | Q9C6U1 | 8e-80 | MYB83_ARATH; Transcription factor MYB83 | ||||
| TrEMBL | A0A2H3YPJ0 | 0.0 | A0A2H3YPJ0_PHODC; transcription factor MYB83 | ||||
| STRING | XP_008801940.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP9821 | 35 | 45 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G08500.1 | 5e-76 | myb domain protein 83 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



