 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Eucgr.I01791.1.p | ||||||||
| Common Name | EUGRSUZ_I01791, LOC104419258 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 211aa MW: 24296.4 Da PI: 7.9829 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 62.3 | 9.7e-20 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg WT eEd++l d+v+++G g+W++Iar+ g++R++k+c++rw +yl
Eucgr.I01791.1.p 14 RGLWTVEEDKILMDYVTEHGRGQWNSIARRTGLRRCGKSCRLRWMNYL 61
788*******************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 61.8 | 1.4e-19 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
r+ +T+eE++l+v+++++lG++ W++Ia++++ gRt++q+k++w++yl
Eucgr.I01791.1.p 67 RDHFTEEEEDLIVRLHNLLGNR-WSLIAKRVP-GRTDNQVKNYWNTYL 112
6789******************.*********.*************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 18.354 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.08E-31 | 12 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.7E-14 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 5.0E-17 | 14 | 61 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.0E-24 | 15 | 68 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.63E-11 | 17 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 28.184 | 62 | 116 | IPR017930 | Myb domain |
| SMART | SM00717 | 2.2E-18 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.1E-18 | 67 | 112 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-26 | 69 | 116 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 7.94E-14 | 70 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010053 | Biological Process | root epidermal cell differentiation | ||||
| GO:0010091 | Biological Process | trichome branching | ||||
| GO:0048629 | Biological Process | trichome patterning | ||||
| GO:0090377 | Biological Process | seed trichome initiation | ||||
| GO:0090378 | Biological Process | seed trichome elongation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 211 aa Download sequence Send to blast |
MTKTEGGGGS QLKRGLWTVE EDKILMDYVT EHGRGQWNSI ARRTGLRRCG KSCRLRWMNY 60 LSPDIKRDHF TEEEEDLIVR LHNLLGNRWS LIAKRVPGRT DNQVKNYWNT YLSKKLGIKK 120 QICQVGRTKS RILHISESST HKDSISSCAG NSRSSEMNVD KSPESQEMLD LPELTSREEY 180 STPSFWDFES DLEMNLSMME FLDGHSLPSY * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 2e-29 | 8 | 116 | 1 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator, when associated with BHLH2/EGL3/MYC146 or BHLH12/MYC1. Regulates the epidermal cell fate specification. Mediates the formation of columellae and accumulation of mucilages on seed coats. Controls the elongation of epidermal cells positively in roots but negatively in stems, leading to the promotion of primary roots elongation and repression of leaves and stems elongation, respectively. Ovoids ectopic root-hair formation, probably by inducing GL2 in roots. Controls trichome initiation and branching. {ECO:0000269|PubMed:11437443, ECO:0000269|PubMed:15361138, ECO:0000269|PubMed:15668208, ECO:0000269|PubMed:15728674}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00538 | DAP | Transfer from AT5G40330 | Download |
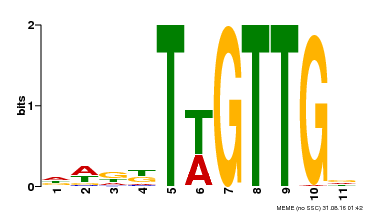
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Eucgr.I01791.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010029175.1 | 1e-156 | PREDICTED: transcription factor MYB23 isoform X2 | ||||
| Swissprot | Q96276 | 2e-71 | MYB23_ARATH; Transcription factor MYB23 | ||||
| TrEMBL | A0A059APY4 | 1e-154 | A0A059APY4_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010029175.1 | 1e-155 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4 | 28 | 2646 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G40330.1 | 9e-74 | myb domain protein 23 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Eucgr.I01791.1.p |
| Entrez Gene | 104419258 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



