 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Eucgr.H00158.1.p | ||||||||
| Common Name | EUGRSUZ_H00158 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 267aa MW: 29799.1 Da PI: 4.8002 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 51.7 | 2.1e-16 | 5 | 52 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg+WT++Ed+ l+ ++++ G g+W++ ++ g+ R++k+c++rw +yl
Eucgr.H00158.1.p 5 RGPWTPDEDQVLISYIQRNGHGNWRALPKQAGLLRCGKSCRLRWINYL 52
89******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 51.1 | 3.1e-16 | 58 | 103 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++ eE+e +++++++lG++ W++Ia++++ gRt++++k+ w+++l
Eucgr.H00158.1.p 58 RGNFNSEEEETIIKLHEMLGNR-WSAIAARLP-GRTDNEIKNVWHTHL 103
89********************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 6.79E-30 | 1 | 99 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 18.114 | 1 | 52 | IPR017930 | Myb domain |
| SMART | SM00717 | 3.3E-12 | 4 | 54 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.5E-15 | 5 | 52 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.5E-24 | 6 | 59 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 4.86E-10 | 7 | 52 | No hit | No description |
| PROSITE profile | PS51294 | 24.699 | 53 | 107 | IPR017930 | Myb domain |
| SMART | SM00717 | 3.6E-14 | 57 | 105 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 9.6E-15 | 58 | 103 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 9.60E-11 | 60 | 103 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 5.1E-26 | 60 | 108 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 267 aa Download sequence Send to blast |
MGLKRGPWTP DEDQVLISYI QRNGHGNWRA LPKQAGLLRC GKSCRLRWIN YLRPDIKRGN 60 FNSEEEETII KLHEMLGNRW SAIAARLPGR TDNEIKNVWH THLKKRLKQN QVTPATTTTN 120 NKSFNLDSSE PTIPRLLESS VHAPMSPQPS SSEISSVTDA STSTGIGEAN NIEIKGEDME 180 YSMESFPLID ESFWSDNPGF LPDEYSSDSS GLSCFNVDTQ TQYSVAPMAD NVQGWPSDGV 240 SHVDNGMDFW YDLFIRAGGA EELPAF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 9e-26 | 3 | 107 | 5 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 102 | 108 | LKKRLKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in cold-regulation of CBF genes and in the development of freezing tolerance. May be part of a complex network of transcription factors controlling the expression of CBF genes and other genes in response to cold stress. Binds to the MYB recognition sequences in the promoters of CBF1, CBF2 and CBF3 genes (PubMed:17015446). Involved in drought and salt tolerance. May enhance expression levels of genes involved in abscisic acid (ABA) biosynthesis and signaling, as well as those encoding stress-protective proteins (PubMed:19161942). {ECO:0000269|PubMed:17015446, ECO:0000269|PubMed:19161942}. | |||||
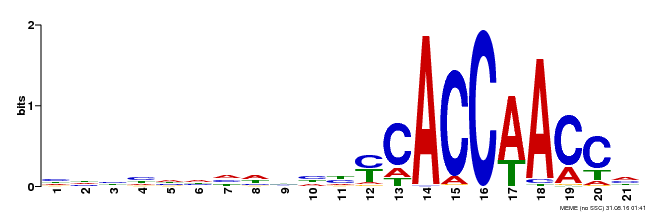
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00375 | DAP | Transfer from AT3G23250 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Eucgr.H00158.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by abscisic acid (ABA) and drought stress (PubMed:19161942). Induced by salt stress (PubMed:19161942). {ECO:0000269|PubMed:19161942}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010069126.1 | 0.0 | PREDICTED: myb-related protein Myb4 | ||||
| Swissprot | Q9LTC4 | 5e-78 | MYB15_ARATH; Transcription factor MYB15 | ||||
| TrEMBL | A0A059ATU1 | 0.0 | A0A059ATU1_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010069126.1 | 0.0 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4 | 28 | 2646 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G23250.1 | 8e-76 | myb domain protein 15 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Eucgr.H00158.1.p |



