 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Eucgr.E01414.1.p | ||||||||
| Common Name | EUGRSUZ_E01414, LOC104444342 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | LBD | ||||||||
| Protein Properties | Length: 245aa MW: 24872.2 Da PI: 7.7391 | ||||||||
| Description | LBD family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF260 | 129.7 | 1.2e-40 | 33 | 133 | 1 | 100 |
DUF260 1 aCaaCkvlrrkCakdCvlapyfpaeq.pkkfanvhklFGasnvlkllkalpeeeredamsslvyeAearardPvyGavgvilklqqqleqlka 92
+C aCk+lrrkC+++C++apyf +eq +++fa+vhk+FGasnv+kll ++p ++r da+ +++yeA+ar+rdPvyG+v++i++lqqq+ +l+a
Eucgr.E01414.1.p 33 PCGACKFLRRKCVPGCIFAPYFDSEQgAAHFAAVHKVFGASNVSKLLLHIPVHRRLDAVVTICYEAQARLRDPVYGCVAHIFALQQQVVNLQA 125
7***********************9989***************************************************************** PP
DUF260 93 elallkee 100
el++l+++
Eucgr.E01414.1.p 126 ELSYLQAH 133
***99986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50891 | 22.999 | 32 | 134 | IPR004883 | Lateral organ boundaries, LOB |
| Pfam | PF03195 | 9.8E-40 | 33 | 131 | IPR004883 | Lateral organ boundaries, LOB |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0010311 | Biological Process | lateral root formation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 245 aa Download sequence Send to blast |
MSSNPSSSGG GGGGGGGSGS GGGSGSGSGS GGPCGACKFL RRKCVPGCIF APYFDSEQGA 60 AHFAAVHKVF GASNVSKLLL HIPVHRRLDA VVTICYEAQA RLRDPVYGCV AHIFALQQQV 120 VNLQAELSYL QAHLTTLELP VPLPPPPPPP AMGPQLSIAD LPSASSIPAT YDLSSLFDPM 180 AQAAWMVPQR QIDFRQFEGS CGGGGSSAAG GGDLNALARE LLRRQGSPPP GCIDAMSSPS 240 SLSK* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5ly0_A | 6e-39 | 29 | 134 | 7 | 111 | LOB family transfactor Ramosa2.1 |
| 5ly0_B | 6e-39 | 29 | 134 | 7 | 111 | LOB family transfactor Ramosa2.1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 7 | 17 | SGGGGGGGGGS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the positive regulation of tracheary element (TE) differentiation. Involved in a positive feedback loop that maintains or promotes NAC030/VND7 expression that regulates TE differentiation-related genes (PubMed:19088331). Functions in the initiation and emergence of lateral roots, in conjunction with LBD16, downstream of ARF7 and ARF19 (PubMed:19717544, PubMed:23749813). Transcriptional activator that directly regulates EXPA14, a gene encoding a cell wall-loosening factor that promotes lateral root emergence. Activates EXPA14 by directly binding to a specific region of its promoter (PubMed:22974309). Transcriptional activator that directly regulates EXPA17, a gene encoding a cell wall-loosening factor that promotes lateral root emergence (PubMed:23872272). Acts downstream of the auxin influx carriers AUX1 and LAX1 in the regulation of lateral root initiation and development (PubMed:26059335). {ECO:0000269|PubMed:19088331, ECO:0000269|PubMed:19717544, ECO:0000269|PubMed:22974309, ECO:0000269|PubMed:23749813, ECO:0000269|PubMed:23872272, ECO:0000269|PubMed:26059335}. | |||||
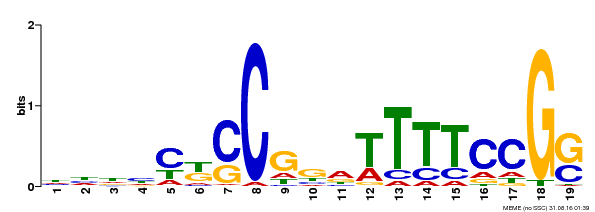
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00314 | DAP | Transfer from AT2G45420 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Eucgr.E01414.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. {ECO:0000269|PubMed:15659631, ECO:0000269|PubMed:19717544, ECO:0000269|PubMed:23749813}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC118672 | 4e-82 | AC118672.3 Genomic sequence for Oryza sativa, Nipponbare strain, clone OSJNBb0078P24, from chromosome 3, complete sequence. | |||
| GenBank | AC226706 | 4e-82 | AC226706.2 Oryza minuta clone OM__Ba0084M12, complete sequence. | |||
| GenBank | AP014959 | 4e-82 | AP014959.1 Oryza sativa Japonica Group DNA, chromosome 3, cultivar: Nipponbare, complete sequence. | |||
| GenBank | CP012611 | 4e-82 | CP012611.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 3 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010056297.1 | 1e-175 | PREDICTED: LOB domain-containing protein 18 | ||||
| Swissprot | O22131 | 1e-83 | LBD18_ARATH; LOB domain-containing protein 18 | ||||
| TrEMBL | A0A059C3F2 | 1e-173 | A0A059C3F2_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010056297.1 | 1e-174 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM108 | 28 | 359 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G45420.1 | 1e-81 | LOB domain-containing protein 18 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Eucgr.E01414.1.p |
| Entrez Gene | 104444342 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



