 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Eucgr.D02105.1.p | ||||||||
| Common Name | EUGRSUZ_D02105, LOC104441939 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 288aa MW: 31824.7 Da PI: 7.4113 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58.7 | 9.8e-19 | 129 | 183 | 2 | 56 |
T--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 2 rkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
rk+ +++k+q +Lee F+++++++ +++ LAk+lgL rqV vWFqNrRa+ k
Eucgr.D02105.1.p 129 RKKLRLSKDQSAVLEESFREHNTLNPKQKLALAKQLGLRPRQVEVWFQNRRARTK 183
788899***********************************************98 PP
| |||||||
| 2 | HD-ZIP_I/II | 128.4 | 3e-41 | 129 | 218 | 1 | 91 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreel 91
+kk+rlsk+q+++LEesF+e+++L+p++K +la++Lgl+prqv+vWFqnrRARtk+kq+E d+e+Lkr++++l+een+rL+kev+eLr +l
Eucgr.D02105.1.p 129 RKKLRLSKDQSAVLEESFREHNTLNPKQKLALAKQLGLRPRQVEVWFQNRRARTKLKQTEIDCEFLKRCCENLTEENRRLQKEVQELR-AL 218
69*************************************************************************************9.55 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF04618 | 2.1E-26 | 2 | 101 | IPR006712 | HD-ZIP protein, N-terminal |
| Gene3D | G3DSA:1.10.10.60 | 2.5E-18 | 104 | 186 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.42E-18 | 119 | 186 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.572 | 125 | 185 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 3.4E-16 | 127 | 189 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 3.5E-16 | 129 | 183 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 6.09E-16 | 129 | 186 | No hit | No description |
| PRINTS | PR00031 | 1.9E-5 | 156 | 165 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 160 | 183 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.9E-5 | 165 | 181 | IPR000047 | Helix-turn-helix motif |
| SMART | SM00340 | 6.6E-27 | 185 | 228 | IPR003106 | Leucine zipper, homeobox-associated |
| Pfam | PF02183 | 1.2E-10 | 185 | 219 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene3D | G3DSA:1.20.5.170 | 6.8E-4 | 189 | 218 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009641 | Biological Process | shade avoidance | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 288 aa Download sequence Send to blast |
MMVEREDLGL SLSLSFSDSS RPSQLGASPF GFNLYKPSHR DCETFASLDR ISEADARPSL 60 RGIDVNRPPP SAADCEEQEE AGVSSPNSTI SSVSGKRGER EMVSGGEDNE AERDCSRGGS 120 DEEDGENSRK KLRLSKDQSA VLEESFREHN TLNPKQKLAL AKQLGLRPRQ VEVWFQNRRA 180 RTKLKQTEID CEFLKRCCEN LTEENRRLQK EVQELRALKL SPQFYMHMPP PTTLTVCPNC 240 ERVGAAAPPL PSAGGGGRPA HHREPVPMIP WAARPGPVSH GALRPRT* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 127 | 133 | SRKKLRL |
| 2 | 177 | 185 | RRARTKLKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that plays a role in auxin-mediated morphogenesis. Negatively regulates lateral root elongation. {ECO:0000269|PubMed:12492842}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00550 | DAP | Transfer from AT5G47370 | Download |
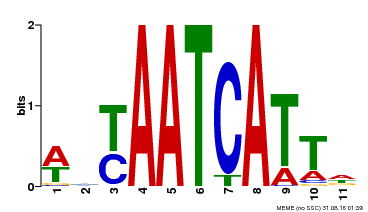
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Eucgr.D02105.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By indole-3-acetic acid (IAA). {ECO:0000269|PubMed:12492842}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010053507.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein HAT4 | ||||
| Swissprot | P46601 | 1e-100 | HAT2_ARATH; Homeobox-leucine zipper protein HAT2 | ||||
| TrEMBL | A0A059CHF5 | 0.0 | A0A059CHF5_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010053507.1 | 0.0 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1563 | 26 | 90 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16780.1 | 1e-82 | homeobox protein 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Eucgr.D02105.1.p |
| Entrez Gene | 104441939 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



