 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Eucgr.B03551.1.p | ||||||||
| Common Name | EUGRSUZ_B03551, LOC104433724 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 788aa MW: 88090.6 Da PI: 7.0811 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 74.6 | 1.2e-23 | 131 | 232 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-S CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldg 86
f+k+lt sd++++g +++ +++a+e+ ++ + +++l+ +d++g++W++++i+r++++r++l++GW+ Fv++++L +gD ++F +
Eucgr.B03551.1.p 131 FCKTLTASDTSTHGGFSVLRRHADEClpqldmtKQ--PPTQELVAKDLHGNEWRFRHIFRGQPRRHLLQSGWSLFVSSKKLVAGDAFIFL--R 219
99****************************97433..34459************************************************..4 PP
SSEE..EEEEE-S CS
B3 87 rsefelvvkvfrk 99
+++el+v+v+r+
Eucgr.B03551.1.p 220 GENGELRVGVRRA 232
49999*****996 PP
| |||||||
| 2 | Auxin_resp | 114 | 1.3e-37 | 257 | 339 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha+st+++F+v+Y+Pr+s++eF+++++k+ ++ k +s+GmRf+m fe+e+++e+r+sGtv+g +d+dp rWp+SkWr+Lk
Eucgr.B03551.1.p 257 AWHAISTGTMFTVYYKPRTSPAEFIIPFDKYIESAKFDYSIGMRFRMTFEGEEAPEQRFSGTVIGSEDVDPPRWPGSKWRCLK 339
89*******************************************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 2.88E-39 | 123 | 260 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 7.0E-40 | 124 | 245 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 6.20E-21 | 129 | 231 | No hit | No description |
| SMART | SM01019 | 1.4E-22 | 131 | 233 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 11.956 | 131 | 233 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.3E-21 | 131 | 232 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 2.5E-35 | 257 | 339 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 4.6E-11 | 663 | 765 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 2.8E-11 | 671 | 750 | No hit | No description |
| PROSITE profile | PS51745 | 23.027 | 674 | 767 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 788 aa Download sequence Send to blast |
MASHEVSVVG SDRRERERTG CEDALYKELW HACAGPLVTV PREGELVYYF PQGHIEQIEA 60 SMNQVADRQI PFYNLPSKIL CRVINVQLRA EPETDELFAQ VTLLPVPNQD ETAVEKETGH 120 PLPPRPRVHS FCKTLTASDT STHGGFSVLR RHADECLPQL DMTKQPPTQE LVAKDLHGNE 180 WRFRHIFRGQ PRRHLLQSGW SLFVSSKKLV AGDAFIFLRG ENGELRVGVR RAMRQLNNVP 240 SSIMPSHSMH IGVLATAWHA ISTGTMFTVY YKPRTSPAEF IIPFDKYIES AKFDYSIGMR 300 FRMTFEGEEA PEQRFSGTVI GSEDVDPPRW PGSKWRCLKV RWDEITSIHR PDRVSPWNIE 360 PAVTAPDNLP ASRPKRPRAS MSSSTDSSVR MREGPLGNGT DPPADIGFSK ALQGQEFSTL 420 KGSLTLNNDM ENAEMPDAWY EFPGDNRKSM HFGQRKCRPN TWVPQLVQEA TLINLVPDSG 480 TFHHRHGIFS SFLRRDSEIT KRAKKQNSAE ETKTDLIPSP GSVRQGPSLN MLGSGIVASL 540 ENESRQQQSL GNVKGDTKRE MNSGNCFLPM HSPSLAQQEL IKANGNGSCK LFGISLMSGT 600 VATESDNGHV HGSQKPVLQS NLVDYSEPNV LGSPIKCLKI APSPIAYGEQ GRAFPSSEQP 660 VRDVEGKHQA VSTRSCIKVH KQDIPVGRSV DLSRFHGYSE LITELDEIFD FNGDLVAPSK 720 EWLVVFTDDE GDMMLVGDDP WEEFCGMVHK IFIYSREEVK RMAPRPLVPK SEEISPKTSH 780 KRYSGGI* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-171 | 23 | 361 | 17 | 354 | Auxin response factor 1 |
| 4ldw_A | 1e-171 | 23 | 361 | 17 | 354 | Auxin response factor 1 |
| 4ldw_B | 1e-171 | 23 | 361 | 17 | 354 | Auxin response factor 1 |
| 4ldx_A | 1e-171 | 23 | 361 | 17 | 354 | Auxin response factor 1 |
| 4ldx_B | 1e-171 | 23 | 361 | 17 | 354 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs) (By similarity). Could act as transcriptional activator or repressor (PubMed:26959229, PubMed:26716451, PubMed:27645097, PubMed:28167023). Involved in the control of fruit ripening process (PubMed:26959229, PubMed:26716451, PubMed:28167023). Regulates expression of a number of ripening regulators, transcription factors, and ethylene biosynthesis and signaling components (PubMed:26959229, PubMed:26716451). May act as a transcriptional repressor of auxin-responsive genes (PubMed:26716451). Regulates vegetative growth, lateral root formation and flower organ senescence, possibly partially by regulating gene expression of auxin and ethylene response factor (ERF) genes (PubMed:28167023). Plays a negative role in axillary shoot meristem formation (PubMed:27645097). {ECO:0000255|RuleBase:RU004561, ECO:0000269|PubMed:26716451, ECO:0000269|PubMed:26959229, ECO:0000269|PubMed:27645097, ECO:0000269|PubMed:28167023}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
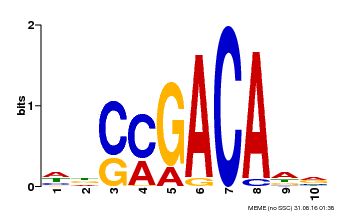
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Eucgr.B03551.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By tomato fruit ripening. Expression in tomato fruits is up-regulated by exogenous application of ethylene and down-regulated by inhibition of ethylene receptors with 1-methylcyclopropene (1-MCP) (PubMed:26959229, PubMed:26716451). Up-regulated by plant hormone abscisic acid (ABA) in tomato fruits (PubMed:26959229). Expression is not up-regulated by plant hormone auxin in tomato fruits (PubMed:26716451). Up-regulated by auxin and gibberellic acid (GA), and down-regulated by ethylene in seedlings (PubMed:28167023). Down-regulated during decapitation-induced axillary shoot development and by excision of immature leaves. Down-regulation is inhibited by the application of auxin on the cut surface (PubMed:27645097). {ECO:0000269|PubMed:26716451, ECO:0000269|PubMed:26959229, ECO:0000269|PubMed:27645097, ECO:0000269|PubMed:28167023}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010044876.1 | 0.0 | PREDICTED: auxin response factor 2 | ||||
| Swissprot | Q2LAJ3 | 0.0 | ARF2A_SOLLC; Auxin response factor 2A | ||||
| TrEMBL | A0A059D8V3 | 0.0 | A0A059D8V3_EUCGR; Auxin response factor | ||||
| STRING | XP_010044876.1 | 0.0 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5348 | 26 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Eucgr.B03551.1.p |
| Entrez Gene | 104433724 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



