 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Eucgr.B03537.1.p | ||||||||
| Common Name | EUGRSUZ_B03537, LOC104433712 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 254aa MW: 28785.7 Da PI: 9.6554 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 165.8 | 1.6e-51 | 14 | 142 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpkkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkevl 93
lppGfrFhPtdeelv +yL++kv + +l++ ++i+ev+++k++PwdLp +++ ++e yfFs+r+ ky++g+r+nrat sgyWkatg d++++
Eucgr.B03537.1.p 14 LPPGFRFHPTDEELVLQYLRRKVYSFPLPA-SIIPEVEVCKSDPWDLPGQCDL-DQERYFFSTREAKYPNGNRSNRATVSGYWKATGIDRQIV 104
79****************************.89***************76655.789************************************ PP
NAM 94 sk...kgelvglkktLvfykgrapkgektdWvmheyrl 128
s+ ++++vg+kktLvfy+g+ p+g+ktdW+mheyrl
Eucgr.B03537.1.p 105 SSrgsQSQVVGMKKTLVFYRGKPPSGSKTDWIMHEYRL 142
*97777778***************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 1.29E-58 | 9 | 175 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 57.28 | 14 | 175 | IPR003441 | NAC domain |
| Pfam | PF02365 | 1.3E-26 | 15 | 142 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 254 aa Download sequence Send to blast |
MDKLNFVKNG VLRLPPGFRF HPTDEELVLQ YLRRKVYSFP LPASIIPEVE VCKSDPWDLP 60 GQCDLDQERY FFSTREAKYP NGNRSNRATV SGYWKATGID RQIVSSRGSQ SQVVGMKKTL 120 VFYRGKPPSG SKTDWIMHEY RLVNPEASVP PKIMNLNQTP VEPMESWVLC RIFLKRRSSK 180 NEGETPVPCQ NLKEAPKSRK NKPVFYDFMT RDLNLAPSSS SSGSSGVTEV SSHNNTDEHE 240 ESSCNSLRKA LKH* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 3e-51 | 12 | 180 | 15 | 170 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 3e-51 | 12 | 180 | 15 | 170 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 3e-51 | 12 | 180 | 15 | 170 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 3e-51 | 12 | 180 | 15 | 170 | NO APICAL MERISTEM PROTEIN |
| 4dul_A | 3e-51 | 12 | 180 | 15 | 170 | NAC domain-containing protein 19 |
| 4dul_B | 3e-51 | 12 | 180 | 15 | 170 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional repressor that negatively regulates the expression of genes involved in xylem vessel formation. Represses the transcriptional activation activity of NAC030/VND7, which regulates protoxylem vessel differentiation by promoting immature xylem vessel-specific genes expression (PubMed:20388856). Transcriptional activator that regulates the COLD-REGULATED (COR15A and COR15B) and RESPONSIVE TO DEHYDRATION (LTI78/RD29A and LTI65/RD29B) genes by binding directly to their promoters. Mediates signaling crosstalk between salt stress response and leaf aging process (PubMed:21673078). May play a role in DNA replication of mungbean yellow mosaic virus (PubMed:24442717). {ECO:0000269|PubMed:20388856, ECO:0000269|PubMed:21673078, ECO:0000269|PubMed:24442717}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00507 | DAP | Transfer from AT5G13180 | Download |
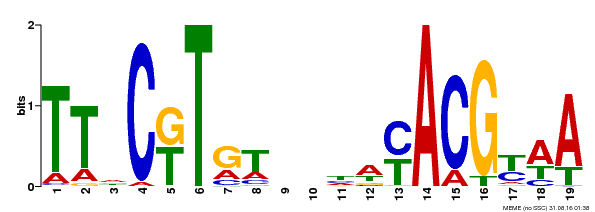
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Eucgr.B03537.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By abscisic acid (ABA) and salt stress. {ECO:0000269|PubMed:21673078}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010044858.1 | 0.0 | PREDICTED: NAC domain-containing protein 83 isoform X1 | ||||
| Swissprot | Q9FY93 | 3e-94 | NAC83_ARATH; NAC domain-containing protein 83 | ||||
| TrEMBL | A0A059D8T7 | 0.0 | A0A059D8T7_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010044858.1 | 0.0 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1806 | 27 | 86 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G13180.1 | 2e-86 | NAC domain containing protein 83 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Eucgr.B03537.1.p |
| Entrez Gene | 104433712 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



