 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EcS754042.10 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 334aa MW: 37471.3 Da PI: 9.7 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 43 | 1.1e-13 | 59 | 107 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s+ykGV + +grW A+I+d r+r+++g+f ++eAa a++ a+k+++g
EcS754042.10 59 SKYKGVVPQP-NGRWGAQIYDN-----RQRVWIGTFSKEDEAALAYDIAAKRFRG 107
89****9888.8*********4.....6**********99*************98 PP
| |||||||
| 2 | B3 | 96.2 | 2.2e-30 | 192 | 297 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsef 90
f+k+ t+sdv+k++r+v+pk++ae+h + + +++ l + d+ g++W+++++y+++s++yvlt+GW +Fvk++ L++gD+v F+++ e+
EcS754042.10 192 FQKEVTRSDVGKLNRFVIPKQHAEKHfplrnrtSPTTSKGVLLNFIDSGGKVWRFRYSYWNSSRSYVLTRGWTRFVKEKSLRTGDIVSFHRSAGPEK 288
99**************************99987555568999************************************************9988889 PP
..EEEEE-S CS
B3 91 elvvkvfrk 99
+l++k r+
EcS754042.10 289 QLHIKCMRR 297
899999887 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 1.44E-16 | 59 | 116 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 9.2E-9 | 59 | 107 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 3.69E-24 | 59 | 114 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 3.0E-19 | 60 | 116 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 2.3E-27 | 60 | 121 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.427 | 60 | 115 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 3.4E-37 | 183 | 298 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.57E-29 | 190 | 298 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 3.89E-26 | 191 | 283 | No hit | No description |
| PROSITE profile | PS50863 | 13.987 | 192 | 298 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 2.7E-26 | 192 | 297 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 3.4E-23 | 192 | 298 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 334 aa Download sequence Send to blast |
MDGNSICIDN LSSTSDSNFI PLPSKSLQPQ SLCHLGSGTG VILDCEGAIE AESRKQPSSK 60 YKGVVPQPNG RWGAQIYDNR QRVWIGTFSK EDEAALAYDI AAKRFRGRDA ITNFKQRSDT 120 EDCDLEAAFL NSHSKEEIVD MLRKHAYYNE LEQSKRKYFN ANGSGNRVSM MMEAIRDPSG 180 PDRAAMACHL LFQKEVTRSD VGKLNRFVIP KQHAEKHFPL RNRTSPTTSK GVLLNFIDSG 240 GKVWRFRYSY WNSSRSYVLT RGWTRFVKEK SLRTGDIVSF HRSAGPEKQL HIKCMRRSGL 300 SQAGPIHSVQ MVRLFGVDIF EVSVEDGGGY GGGS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 2e-47 | 191 | 305 | 13 | 127 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional repressor of flowering time on long day plants. Acts directly on FT expression by binding 5'-CAACA-3' and 5'-CACCTG-3 sequences. Functionally redundant with TEM2. {ECO:0000269|PubMed:18718758}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00024 | PBM | Transfer from AT1G25560 | Download |
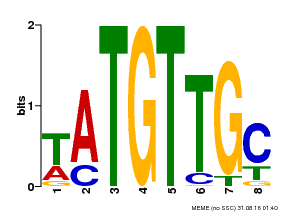
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Expressed with a circadian rhythm showing a peak at dawn. {ECO:0000269|PubMed:18718758}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010046597.2 | 0.0 | PREDICTED: AP2/ERF and B3 domain-containing transcription repressor TEM1-like | ||||
| Swissprot | Q9C6M5 | 1e-121 | RAVL1_ARATH; AP2/ERF and B3 domain-containing transcription repressor TEM1 | ||||
| TrEMBL | A0A058ZSG9 | 0.0 | A0A058ZSG9_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010046597.1 | 0.0 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM848 | 27 | 121 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25560.1 | 1e-123 | RAV family protein | ||||



