 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EcC045126.10 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 688aa MW: 75207.3 Da PI: 7.3805 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 37.8 | 4.8e-12 | 322 | 378 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pse.ng.krkrfslgkfgtaeeAakaaiaarkkleg 55
s+y+GV++++++gr++A+++d ++ g rk + g ++ +e+Aa+a++ a++k++g
EcC045126.10 322 SQYRGVTRHRWTGRYEAHLWDnSCKkEGqTRKGRQ-GGYDKEEKAARAYDLAALKYWG 378
78*******************77776664446655.779999**************98 PP
| |||||||
| 2 | AP2 | 45.6 | 1.8e-14 | 423 | 472 | 3 | 55 |
AP2 3 ykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
y+GV+++++ grW A+I + +k +lg+f t eeAa+a++ a+ k++g
EcC045126.10 423 YRGVTRHHQHGRWQARIGRVAG---NKDLYLGTFSTQEEAAEAYDIAAIKFRG 472
9***************988532...5************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 2.26E-19 | 322 | 386 | No hit | No description |
| SuperFamily | SSF54171 | 1.31E-14 | 322 | 388 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 7.2E-9 | 322 | 378 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.2E-12 | 323 | 387 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 17.425 | 323 | 386 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 5.2E-23 | 323 | 392 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 4.5E-6 | 324 | 335 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.57E-17 | 421 | 482 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 6.37E-25 | 421 | 482 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 4.4E-18 | 422 | 480 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 7.5E-34 | 422 | 486 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.177 | 422 | 480 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 3.0E-9 | 423 | 472 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 4.5E-6 | 462 | 482 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007276 | Biological Process | gamete generation | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0042127 | Biological Process | regulation of cell proliferation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 688 aa Download sequence Send to blast |
MKSVSENGGG ASGGNWLGFS LSPHHLKMEV APQAEHHQNH HHHHQYHHQP QQGEPGNYFL 60 SAPPPPPHYS SSGLCYGGGV GDNNNGGYLH SPLSVMPLKS DGSLCIMEAL TRSRPQGLGQ 120 GSTPKLEDFL GGASATVTAT TVPLSLDSLY SYQQSADPEK QSLDLLNEPF RHQQQHQIYY 180 PNELTCPALY HSPLAEEEPK DTHIASCDEN SGFSCLKSWV TRNFSSSSCH VGLDQMSSGM 240 VCHNGGNSSE AVGPMECGDL QNLSLSMSPS STSSCVTVPT RQISPTENEC VGMEMTKKRS 300 SNCHNKQPVH RKSIDTFGQR TSQYRGVTRH RWTGRYEAHL WDNSCKKEGQ TRKGRQGGYD 360 KEEKAARAYD LAALKYWGPS THINFPLENY QKELEEMKNM SRQEYVAHLR RKSSGFSRGA 420 SMYRGVTRHH QHGRWQARIG RVAGNKDLYL GTFSTQEEAA EAYDIAAIKF RGVNAVTNFD 480 ISRYDVERIM ASSTLLAGEL ARRTKETAPN EEVAEYNALT QQKIGSDGTL QLESSGHVTN 540 GSDWKMVLYH SSPQLQASCN QSLDHNSVGC PSGGGNYRNS SFSMALHDLI GMESSVNSSQ 600 PSVDDPAKLG TRHLSNPSSL VTSLSSSREA SPDRAGSAFA NKPPIVSKLV NASPTSASVT 660 SWLPSPAHMR PSAIAMSHLP VFAAWNDT |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Acts as positive regulator of adventitious (crown) root formation by promoting its initiation. Promotes adventitious root initiation through repression of cytokinin signaling by positively regulating the two-component response regulator RR1. Regulated by the auxin response factor and transcriptional activator ARF23/ARF1. {ECO:0000269|PubMed:21481033}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00088 | SELEX | Transfer from AT4G37750 | Download |
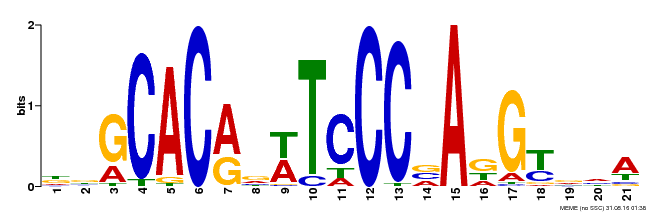
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by auxin. {ECO:0000269|PubMed:21481033}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010061622.1 | 0.0 | PREDICTED: AP2-like ethylene-responsive transcription factor ANT isoform X1 | ||||
| Swissprot | Q84Z02 | 1e-144 | CRL5_ORYSJ; AP2-like ethylene-responsive transcription factor CRL5 | ||||
| TrEMBL | A0A059BRN1 | 0.0 | A0A059BRN1_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010061621.1 | 0.0 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3137 | 28 | 63 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G37750.1 | 1e-122 | AP2 family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



