 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EcC043315.10 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 431aa MW: 46432.4 Da PI: 7.7334 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.5 | 5.4e-32 | 203 | 258 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+p+LH+rFv+a++ LGGs++AtPk+i+elmkv+gLt+++vkSHLQkYRl+
EcC043315.10 203 KARRCWSPDLHRRFVNALQMLGGSQVATPKQIRELMKVDGLTNDEVKSHLQKYRLH 258
68****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 4.66E-17 | 200 | 261 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 13.355 | 200 | 260 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.9E-28 | 201 | 261 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.2E-26 | 203 | 258 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 7.3E-7 | 205 | 256 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009266 | Biological Process | response to temperature stimulus | ||||
| GO:0009740 | Biological Process | gibberellic acid mediated signaling pathway | ||||
| GO:0010452 | Biological Process | histone H3-K36 methylation | ||||
| GO:0048579 | Biological Process | negative regulation of long-day photoperiodism, flowering | ||||
| GO:1903507 | Biological Process | negative regulation of nucleic acid-templated transcription | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 431 aa Download sequence Send to blast |
KRELPLCMQL LTNAVEVSKQ QLQAYRANSG PKPVLEEFMP LKNSSSEGSE KTPNLSEKAS 60 WMISAQLWSQ TSNDSPNKQQ SLAPKETDIG FNVSPKFSLE NKSRNGGAFL PFSKERNGSC 120 PSPSLRALPE LALGSPDKEM EDKKYPEAEN GNGVSCVRRD GCSKIGNGEV GKGTVSSAEV 180 QIPATANASN TNTTAANGQT HRKARRCWSP DLHRRFVNAL QMLGGSQVAT PKQIRELMKV 240 DGLTNDEVKS HLQKYRLHTR RPSPSPQTAG SAAPQLVVLG GIWVPPDYAT AAAAAVAHGN 300 TSPLYGAHPS AHASPHYCTP PGMPQEFYSA AASHHEAVHH HPLHGHHHQI QVYKGTSHPQ 360 PQSSPESGMR GGGGDGQSES IEDGKSESSS WKGDSGDHVN GGDQRNGLIT MRDDNNEESN 420 GSEITLKFYD K |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 9e-14 | 203 | 256 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 9e-14 | 203 | 256 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 9e-14 | 203 | 256 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 9e-14 | 203 | 256 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
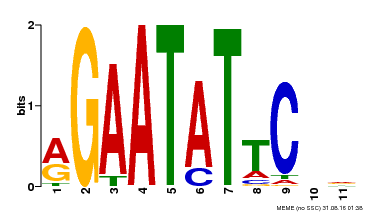
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00261 | DAP | Transfer from AT2G03500 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010049950.1 | 0.0 | PREDICTED: myb family transcription factor EFM isoform X2 | ||||
| TrEMBL | A0A059CX25 | 0.0 | A0A059CX25_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010049949.1 | 0.0 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM7134 | 28 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G03500.1 | 6e-94 | G2-like family protein | ||||



