 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EcC014751.10 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 158aa MW: 18069.8 Da PI: 4.9839 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.3 | 2.9e-32 | 97 | 150 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtp+LH +Fv+ave+LGG+e+AtPk++l+ m+vkgL+++hvkSHLQ+YR+
EcC014751.10 97 PRLRWTPQLHYSFVQAVERLGGQERATPKSVLQQMNVKGLSINHVKSHLQMYRS 150
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 3.1E-30 | 93 | 151 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 11.74 | 93 | 153 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.0E-15 | 95 | 151 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 4.7E-24 | 97 | 152 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 3.7E-10 | 98 | 149 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 158 aa Download sequence Send to blast |
MRETMRSLES TEDHQEKSEE KGEKSSPAVS SQKLSSFDLN EEATFDEHDS PAEDDDLISD 60 EDDNSSIREA EDKPGNRNAE SSETMKAARQ YVRSKMPRLR WTPQLHYSFV QAVERLGGQE 120 RATPKSVLQQ MNVKGLSINH VKSHLQMYRS RKLDEVGQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 9e-21 | 98 | 150 | 4 | 56 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 9e-21 | 98 | 150 | 4 | 56 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 8e-21 | 98 | 150 | 3 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 8e-21 | 98 | 150 | 3 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 8e-21 | 98 | 150 | 3 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 8e-21 | 98 | 150 | 3 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 9e-21 | 98 | 150 | 4 | 56 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 9e-21 | 98 | 150 | 4 | 56 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 9e-21 | 98 | 150 | 4 | 56 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 9e-21 | 98 | 150 | 4 | 56 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 9e-21 | 98 | 150 | 4 | 56 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 9e-21 | 98 | 150 | 4 | 56 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
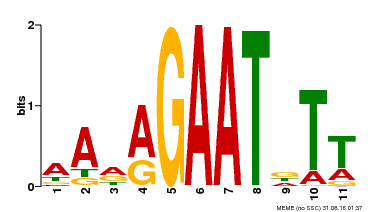
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00297 | DAP | Transfer from AT2G38300 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010057109.1 | 2e-97 | PREDICTED: uncharacterized protein LOC104445014 | ||||
| Refseq | XP_018728583.1 | 2e-97 | PREDICTED: uncharacterized protein LOC104445014 | ||||
| Refseq | XP_018728584.1 | 2e-97 | PREDICTED: uncharacterized protein LOC104445014 | ||||
| Refseq | XP_018728585.1 | 2e-97 | PREDICTED: uncharacterized protein LOC104445014 | ||||
| TrEMBL | A0A059C6L8 | 2e-87 | A0A059C6L8_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010057109.1 | 7e-97 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM363 | 28 | 184 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38300.1 | 6e-36 | G2-like family protein | ||||



