 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EcC010491.30 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 512aa MW: 56449.9 Da PI: 5.085 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 57.4 | 3.3e-18 | 36 | 83 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+ Ed +lvd+v+++G g+W+++ ++ g+ R++k+c++rw ++l
EcC010491.30 36 KGPWTAAEDAILVDYVTKHGEGNWNAVQKHSGLARCGKSCRLRWANHL 83
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 50.1 | 6.2e-16 | 89 | 132 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++++eE+ l++++++++G++ W++ a++++ gRt++++k++w++
EcC010491.30 89 KGAFSPEEERLIIELHAKMGNK-WARMAAELP-GRTDNEIKNYWNT 132
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 16.763 | 31 | 83 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.55E-30 | 33 | 130 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 9.2E-14 | 35 | 85 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.7E-15 | 36 | 83 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.0E-24 | 37 | 90 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.31E-11 | 38 | 83 | No hit | No description |
| PROSITE profile | PS51294 | 25.91 | 84 | 138 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.2E-16 | 88 | 136 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.1E-15 | 89 | 132 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.85E-12 | 91 | 132 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-25 | 91 | 136 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006468 | Biological Process | protein phosphorylation | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0004672 | Molecular Function | protein kinase activity | ||||
| GO:0005524 | Molecular Function | ATP binding | ||||
| GO:0016151 | Molecular Function | nickel cation binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 512 aa Download sequence Send to blast |
MRSTTDGGRD KKVRKIRRHP PSGEEASDGG GIPLKKGPWT AAEDAILVDY VTKHGEGNWN 60 AVQKHSGLAR CGKSCRLRWA NHLRPDLKKG AFSPEEERLI IELHAKMGNK WARMAAELPG 120 RTDNEIKNYW NTRIKRLQRT GMPIYPTEVC LQVSSENQET HNMGNLHTAG EDNCDLSQAD 180 PLEIPEVDFR KLELHLGFSS FWSTLLDVPP CGFGREAMCL SDAYCLPFPS SRSPKRLRGS 240 ETPFPVLDAG SNDSFSSFGQ YADYSSEITP ESFVLSSPYD VNPDSTNQSL LGDLPGSHAF 300 LNGNYPSSVP HLEAMKSELP SLQYTETQQS SCGTPASPRP SLEFIDPLIQ SPPCEHYQSD 360 CHSPRGCGLL EAVIYEAFSK QKSCAQTTVV SGNEACNSSP VQLRTKQEVY AAPDSPLGDN 420 AISVLNHCSP ASRKSFEEPQ SANTMPGFDI KPEEDSKTFP DFEGKQEILD QMHINGPDAL 480 LDLDWCQGKS QPSEDKIGRA DYIGALLGTD FY |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 2e-32 | 31 | 136 | 1 | 106 | B-MYB |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
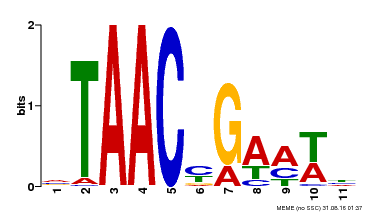
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010045104.1 | 0.0 | PREDICTED: transcription factor GAMYB | ||||
| TrEMBL | A0A059D9S4 | 0.0 | A0A059D9S4_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010045104.1 | 0.0 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3802 | 28 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 3e-86 | myb domain protein 33 | ||||



