 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EcC000253.10 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 354aa MW: 38054.8 Da PI: 9.6435 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 65.3 | 1.2e-20 | 139 | 188 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrd.psengkr.krfslgkfgtaeeAakaaiaarkkleg 55
+y+GVr+++ +g+W+AeIrd + + r++lg+f+tae+Aa+a+++a+++++g
EcC000253.10 139 KYRGVRQRP-WGKWAAEIRDpF-----KaSRVWLGTFDTAEGAARAYDRAALQFRG 188
8********.**********66.....56***********************9998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.730.10 | 7.2E-33 | 138 | 197 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 8.98E-33 | 139 | 198 | No hit | No description |
| PROSITE profile | PS51032 | 25.33 | 139 | 196 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 3.9E-37 | 139 | 202 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.11E-22 | 139 | 197 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 1.0E-13 | 139 | 188 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.3E-11 | 140 | 151 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.3E-11 | 162 | 178 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 354 aa Download sequence Send to blast |
MSIMVSALRR VVSGEVLSGD DQQQQFNLSG GYCSRDGPAS LLGRSSSLDS VGVGQKRGRG 60 DSTLSSEFSV SQDSVALGGF PQAGSSSSSD AREGPAGSTT VETEKAERPA HGTAPKYEYA 120 YEAPTTMAAS RLDPTVRRKY RGVRQRPWGK WAAEIRDPFK ASRVWLGTFD TAEGAARAYD 180 RAALQFRGSK AKLNFPENVR LRQQPAPVAS AAFPSATHFT ISRELPASGN FDATGYDGQA 240 PMQQFPENDF RAYHRDVAAH QERLQGRTMS LYEQMLLSNS SFGSQFQPAF SLSPSSSSSS 300 SSVLARSPNS SLFMPSPPNV TSRSQESRRG SGDGVAVFSQ PPWTDSSHYS SSSG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 1e-22 | 138 | 199 | 4 | 66 | ATERF1 |
| 3gcc_A | 1e-22 | 138 | 199 | 4 | 66 | ATERF1 |
| Search in ModeBase | ||||||
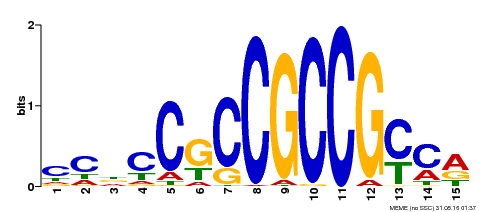
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00289 | DAP | Transfer from AT2G33710 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010027737.1 | 0.0 | PREDICTED: ethylene-responsive transcription factor ABR1 isoform X1 | ||||
| TrEMBL | A0A059AJY4 | 0.0 | A0A059AJY4_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010027737.1 | 0.0 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM10 | 28 | 1650 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G43160.1 | 1e-26 | related to AP2 6 | ||||



