 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Do010386.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Dichantheliinae; Dichanthelium
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 412aa MW: 43756 Da PI: 9.1252 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 130.8 | 5e-41 | 170 | 246 | 1 | 77 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S-- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkq 77
+Cq+egC+adls ak+yhrrhkvCe h+ka+vv+++g++qrfCqqCsrfh l+efDe+krsCr+rLa+hn+rrrk++
Do010386.1 170 RCQAEGCKADLSAAKHYHRRHKVCEYHAKASVVAAAGKQQRFCQQCSRFHVLTEFDEAKRSCRKRLAEHNRRRRKPA 246
6**************************************************************************87 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 5.8E-35 | 163 | 232 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.419 | 168 | 245 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 2.62E-38 | 169 | 248 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 7.9E-31 | 171 | 244 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009554 | Biological Process | megasporogenesis | ||||
| GO:0009556 | Biological Process | microsporogenesis | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 412 aa Download sequence Send to blast |
MMSGRLNVAT SAAGDDFPFV PMQQQHQPPP PYVGFEHGVA GGGGGMQHHL YDGLDFAAAM 60 QFQEAAPHHH QLLTLPSSLG PMAPPPLPMP LQMPMPMPGM PGDVYPALGM VKREGDGASA 120 AAGGRIGLNL GRRTYFSPGD MLAVDRLLMR SRLGGVFGLG FGGAHHQPPR CQAEGCKADL 180 SAAKHYHRRH KVCEYHAKAS VVAAAGKQQR FCQQCSRFHV LTEFDEAKRS CRKRLAEHNR 240 RRRKPATTAS SKDAASPPAK KPSSGSISGS YTTDNKNLTT SKSTMSSNTG SVISCLQDQG 300 SKQLARPTLT LGASPDKQDH QHQQFSTMLH QAQAATGGQH HQEQHFLTSL QVHNHGNGGG 360 GGSGNNNILS CSSVCSSALP SANGEASDQN TTNNNGGGGN GNMHNLFEVD FM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 2e-29 | 171 | 244 | 11 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 227 | 244 | KRSCRKRLAEHNRRRRKP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00090 | SELEX | Transfer from AT1G02065 | Download |
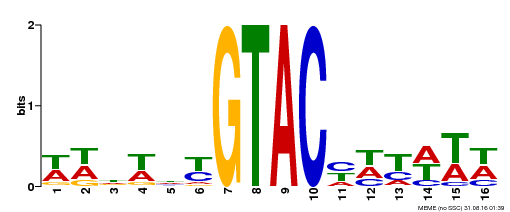
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Do010386.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT033937 | 0.0 | BT033937.1 Zea mays full-length cDNA clone ZM_BFc0033O21 mRNA, complete cds. | |||
| GenBank | BT034298 | 0.0 | BT034298.1 Zea mays full-length cDNA clone ZM_BFc0007A11 mRNA, complete cds. | |||
| GenBank | BT065712 | 0.0 | BT065712.1 Zea mays full-length cDNA clone ZM_BFc0041B17 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004965676.1 | 0.0 | squamosa promoter-binding-like protein 10 | ||||
| Swissprot | A2YFT9 | 1e-111 | SPL10_ORYSI; Squamosa promoter-binding-like protein 10 | ||||
| Swissprot | Q0DAE8 | 1e-111 | SPL10_ORYSJ; Squamosa promoter-binding-like protein 10 | ||||
| TrEMBL | A0A1E5VVH0 | 0.0 | A0A1E5VVH0_9POAL; Squamosa promoter-binding-like protein 10 | ||||
| STRING | Si006559m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2375 | 35 | 93 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G02065.1 | 1e-48 | squamosa promoter binding protein-like 8 | ||||



