 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Do001050.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Dichantheliinae; Dichanthelium
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 552aa MW: 59683.3 Da PI: 4.952 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 52.5 | 1.1e-16 | 43 | 90 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed +lvd+vk++G g+W+++ + g+ R++k+c++rw ++l
Do001050.1 43 KGPWTSAEDAILVDYVKKHGEGNWNAVQKNTGLFRCGKSCRLRWANHL 90
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 46 | 1.2e-14 | 96 | 139 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++ +eE+ l++++++++G++ W++ a++++ gRt++++k++w++
Do001050.1 96 KGAFIPEEERLIIQLHAKMGNK-WARMAAHLP-GRTDNEIKNYWNT 139
678889****************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF001693 | 1.7E-252 | 1 | 552 | IPR016310 | Transcription factor, GAMYB |
| PROSITE profile | PS51294 | 16.625 | 38 | 90 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.17E-30 | 40 | 137 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.1E-14 | 42 | 92 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.1E-14 | 43 | 90 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.5E-22 | 44 | 97 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.68E-11 | 45 | 90 | No hit | No description |
| PROSITE profile | PS51294 | 23.525 | 91 | 145 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.0E-14 | 95 | 143 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 8.2E-14 | 96 | 139 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-24 | 98 | 143 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.04E-11 | 101 | 139 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 552 aa Download sequence Send to blast |
MYRVKSESDC EMMLQDQMDS PVADDGSSGG GSPHRGSGPP LKKGPWTSAE DAILVDYVKK 60 HGEGNWNAVQ KNTGLFRCGK SCRLRWANHL RPNLKKGAFI PEEERLIIQL HAKMGNKWAR 120 MAAHLPGRTD NEIKNYWNTR IKRCQRAGLP IYPASVCNQS SSEDQQLSGD FNCGENISND 180 LLSGNNLYLP DFTSDNFIAN PEALSYAPQL SAVSISNLLG QSFASKNCSF MDQVDQSGIL 240 KQSGCVLPAL SDTIDGVLSS VDQFSNDSEK LKQALGFDYL NEANASSKTI APFGVALSGS 300 HAFFNGNFSA SRPINGPLKM ELPSLQDTES DPNSWLKYTL APAMQPTELV DPYLHSPIAT 360 PSVKSECASP RNSGLLEELL HEAQALGSGK NQQPSVRSSS SSAGTPCETT TVASPEFDLC 420 QEYWDDHHSS FLNECTPFSG YSFTESTPVS AASPDIFHLS KISPQQSPSM GSGEQALEPK 480 HDTAGSPHPE NLRPDALFSG NTADPSTFNN AISILLGNDM NAECKPVLGG GITFGSSSWN 540 NMPHACEIPE FK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 1e-31 | 39 | 143 | 3 | 106 | B-MYB |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator of gibberellin-dependent alpha-amylase expression in aleurone cells. Involved in pollen and floral organs development. May bind to the 5'-TAACAAA-3' box of alpha-amylase promoter. {ECO:0000269|PubMed:9150608}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
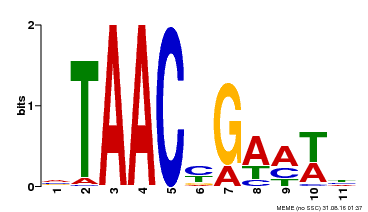
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Do001050.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By gibberellin in aleurone cells. {ECO:0000269|PubMed:9150608}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT033925 | 0.0 | BT033925.1 Zea mays full-length cDNA clone ZM_BFc0044D10 mRNA, complete cds. | |||
| GenBank | BT062692 | 0.0 | BT062692.1 Zea mays full-length cDNA clone ZM_BFb0334D20 mRNA, complete cds. | |||
| GenBank | BT063828 | 0.0 | BT063828.1 Zea mays full-length cDNA clone ZM_BFc0110K24 mRNA, complete cds. | |||
| GenBank | HQ858750 | 0.0 | HQ858750.1 Zea mays clone UT1459 MYBGA transcription factor mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025819835.1 | 0.0 | transcription factor GAMYB-like | ||||
| Swissprot | A2WW87 | 0.0 | GAM1_ORYSI; Transcription factor GAMYB | ||||
| TrEMBL | A0A1E5WEM8 | 0.0 | A0A1E5WEM8_9POAL; Transcription factor | ||||
| STRING | Pavir.Ea03206.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3947 | 36 | 64 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.1 | 3e-80 | myb domain protein 33 | ||||



