 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KZV57631.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Gesneriaceae; Didymocarpoideae; Trichosporeae; Loxocarpinae; Dorcoceras
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 819aa MW: 89341 Da PI: 6.4303 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 64.5 | 1.5e-20 | 126 | 181 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+++l L++rqVk+WFqNrR+++k
KZV57631.1 126 KKRYHRHTPQQIQELETLFKECPHPDEKQRLELSRRLCLETRQVKFWFQNRRTQMK 181
688999***********************************************999 PP
| |||||||
| 2 | START | 203.7 | 7.5e-64 | 335 | 559 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEEEEC CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaetlevis 86
ela++a++elvk+a+ +ep+W ks e +n +e+ +f++ + + +ea+r++g+v+ ++ lve+l+d++ +W+e+++ + +t++vis
KZV57631.1 335 ELALSAMDELVKMAQTDEPLWIKSLeggrEVLNHEEYFNTFTPCIGmkpngFVTEASRETGMVIINSLALVETLMDSN-KWTEMFPciipRTSTTDVIS 432
57899**********************9999**********99988999*9***************************.******************** PP
TT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXH CS
START 87 sg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehvdlkgrlph 178
sg galqlm aelq+lsplvp Rd+ f+R+++q+ +g+w++vdvS+d ++ + + +++lpSg+l+++++ng+skvtwveh++++++++h
KZV57631.1 433 SGmggtrnGALQLMHAELQVLSPLVPvRDVNFLRFCKQHAEGVWAVVDVSIDTVRETAGGPTFSSCRRLPSGCLVQDMPNGYSKVTWVEHAEYDESIIH 531
********************************************************998**************************************** PP
HHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 179 wllrslvksglaegaktwvatlqrqcek 206
+l+r+l+ g+ +ga++wva lqrqce+
KZV57631.1 532 QLYRPLIAAGMGFGAQRWVAMLQRQCEC 559
**************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.04E-20 | 113 | 183 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 4.8E-21 | 118 | 183 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.184 | 123 | 183 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 6.5E-17 | 124 | 187 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 6.16E-18 | 125 | 183 | No hit | No description |
| Pfam | PF00046 | 4.1E-18 | 126 | 181 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 158 | 181 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 43.397 | 326 | 562 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.51E-32 | 328 | 559 | No hit | No description |
| CDD | cd08875 | 4.48E-126 | 330 | 558 | No hit | No description |
| SMART | SM00234 | 1.8E-48 | 335 | 559 | IPR002913 | START domain |
| Pfam | PF01852 | 1.7E-55 | 335 | 559 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.77E-24 | 587 | 812 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 819 aa Download sequence Send to blast |
MNFGDFLDNG GGGGVRIVSD ILYNSNTSTP RSKIKGSMPT GAIAQPLLVS QSLAAKAMFN 60 SPGLSLALQT GMEGQGEVGR IGENFESTNV GGRRSRDEEH ESRSGSDNLD GVSGDELDAA 120 DKPPRKKRYH RHTPQQIQEL ETLFKECPHP DEKQRLELSR RLCLETRQVK FWFQNRRTQM 180 KTQLERHENS LLRQENDKLR AENMSMREMM RNPICTACGG PAIIGEISLE EQHLRIENAR 240 LKDELDRVCT LAGKFIGRPI SSLAASMGLP LPNSNLELGV GNNDFGGLNG LPPALHSGPP 300 HFGSGISPPL PIEQPSKPLN ITPIDRSLER SVYLELALSA MDELVKMAQT DEPLWIKSLE 360 GGREVLNHEE YFNTFTPCIG MKPNGFVTEA SRETGMVIIN SLALVETLMD SNKWTEMFPC 420 IIPRTSTTDV ISSGMGGTRN GALQLMHAEL QVLSPLVPVR DVNFLRFCKQ HAEGVWAVVD 480 VSIDTVRETA GGPTFSSCRR LPSGCLVQDM PNGYSKVTWV EHAEYDESII HQLYRPLIAA 540 GMGFGAQRWV AMLQRQCECL AILMSSSAPA RDHTAITAGG RKSMLKLAQR MTNNFCAGVC 600 ASTVHKWNKL NAGTVDEDVR VMTRKSVDDP GEPPGIVVSA ATSVWLPVSP QRLFDFLRDE 660 RLRSEWDILS NGGPMQEMAH IAKGQDHGNC VSLLRASAMN SNQSSMLILQ ETCIDAAGSL 720 VVYAPVDIPA MHVVMNGGDS AYVALLPSGF VIVPDGPGSR GPASNGMAHR VSGSLLTVAF 780 QILVNSLPTA KLTVESVETV NNLITCTVQK IKAALHCES |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
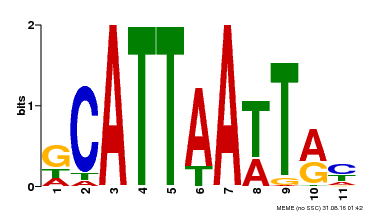
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KZV57631.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010661562.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A2G9HWV6 | 0.0 | A0A2G9HWV6_9LAMI; Transcription factor DLX | ||||
| STRING | VIT_15s0048g02000.t01 | 0.0 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1626 | 24 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



