 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KZV48910.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Gesneriaceae; Didymocarpoideae; Trichosporeae; Loxocarpinae; Dorcoceras
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 263aa MW: 30399.1 Da PI: 8.1065 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 53.2 | 6.9e-17 | 11 | 58 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+W++eEd ll ++v+q+G g+W++++ + g++R++k+c++rw++yl
KZV48910.1 11 KGAWSKEEDALLRKCVQQFGVGRWHLVPLRSGLNRCRKSCRLRWLNYL 58
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 46.3 | 9.6e-15 | 64 | 107 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
rg+ T +E +ll++++k+lG++ W++Ia +++ gRt++++k++w++
KZV48910.1 64 RGNLTRDEIDLLIRLHKLLGNR-WSLIAGRIP-GRTANDLKNFWNT 107
7899******************.*********.***********97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 23.901 | 6 | 62 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.74E-27 | 8 | 105 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 4.3E-15 | 10 | 60 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.3E-15 | 11 | 58 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.5E-20 | 12 | 65 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.76E-10 | 13 | 58 | No hit | No description |
| SMART | SM00717 | 3.8E-13 | 63 | 111 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 17.325 | 63 | 113 | IPR017930 | Myb domain |
| Pfam | PF00249 | 1.2E-12 | 64 | 107 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.1E-20 | 66 | 112 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.72E-9 | 68 | 107 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0031540 | Biological Process | regulation of anthocyanin biosynthetic process | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 263 aa Download sequence Send to blast |
MERAKSLGVR KGAWSKEEDA LLRKCVQQFG VGRWHLVPLR SGLNRCRKSC RLRWLNYLRP 60 DIRRGNLTRD EIDLLIRLHK LLGNRWSLIA GRIPGRTAND LKNFWNTRME KKSREKVQPI 120 QKAITKTSIL RPRPRNFSKL MVHDHHGTRL ATDDFHQSTS DNSTNDNDNK LLPSSSQGSD 180 EDIGWWSNLL DASEDQLEKD ITFADGKGEE EGLWMGQKNP AGLLNDWADH AIPANEFADD 240 DFNFEDDVWE RLGFGNYDAI LQY |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gv2_A | 5e-22 | 11 | 112 | 4 | 104 | MYB PROTO-ONCOGENE PROTEIN |
| 1h88_C | 2e-21 | 11 | 112 | 58 | 158 | MYB PROTO-ONCOGENE PROTEIN |
| 1h89_C | 2e-21 | 11 | 112 | 58 | 158 | MYB PROTO-ONCOGENE PROTEIN |
| 1mse_C | 7e-22 | 11 | 112 | 4 | 104 | C-Myb DNA-Binding Domain |
| 1msf_C | 7e-22 | 11 | 112 | 4 | 104 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator, when associated with BHLH002/EGL3/MYC146, BHLH012/MYC1, or BHLH042/TT8. {ECO:0000269|PubMed:15361138}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00213 | DAP | Transfer from AT1G66370 | Download |
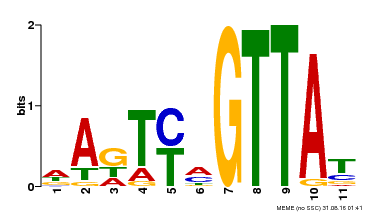
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KZV48910.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011073629.1 | 5e-72 | transcription factor MYB114 | ||||
| Swissprot | Q9FNV9 | 5e-60 | MY113_ARATH; Transcription factor MYB113 | ||||
| TrEMBL | A0A2Z7CPC7 | 0.0 | A0A2Z7CPC7_9LAMI; Uncharacterized protein | ||||
| STRING | XP_009627993.1 | 3e-66 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA12 | 24 | 2154 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G66370.1 | 4e-61 | myb domain protein 113 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



