 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KZV48886.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Gesneriaceae; Didymocarpoideae; Trichosporeae; Loxocarpinae; Dorcoceras
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 496aa MW: 55648.9 Da PI: 6.4239 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 21.1 | 8.2e-07 | 257 | 278 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
C++Cgk F+r nL+ H+r H
KZV48886.1 257 FCTICGKGFKRDANLRMHMRGH 278
6*******************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 2.08E-5 | 254 | 281 | No hit | No description |
| PROSITE profile | PS50157 | 12.03 | 256 | 283 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0026 | 256 | 278 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 7.6E-6 | 257 | 307 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 258 | 278 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 56 | 305 | 338 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 20 | 343 | 365 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010044 | Biological Process | response to aluminum ion | ||||
| GO:0010447 | Biological Process | response to acidic pH | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 496 aa Download sequence Send to blast |
MDHEKRMHAD KWTEPSSSRD THIPTNNQSF SNFDPAQQKW EESSLLDYGM RIDSRNSFAC 60 DTDNQNQTSD PLGNNTGNAT EPINIQEWDP RTMLNNLCFL EKKIHQLQDL IHSFVGRKGQ 120 SSGRPDELLV QQQQLVTVDL TSIIVQLIST AGSLLPSLNH NLSSVNPINQ LGHFSSVANP 180 SNGAPRNIFP KNTNLNKVED HSNLIDTTGL CGTELSNMED QENKSDDDAE EGENLPPGSY 240 EILQLEKEEI LAPHTHFCTI CGKGFKRDAN LRMHMRGHGD EYKTPAALAK PSKESTSEQI 300 LVKRYSCPYI GCKRNRDHKK FEPLKTILCV KNHYKRTHCD KTYTCSRCNA KKFSVLADLK 360 THEKHCGRDK WLCSCGTTFS RKDKLFGHIA LFQGHTPAIP VEDSKGLSTP ANEGQGNEAT 420 SKVEQINFDN QLNARTPVSF QNLVDVKGLA GEDLANFFSP FETNGISGFH EFPRPPFEDS 480 ESSFSFLLSD YTQKNG |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. Together with STOP2, plays a critical role in tolerance to major stress factors in acid soils such as proton H(+) and aluminum ion Al(3+). Required for the expression of genes in response to acidic stress (e.g. ALMT1 and MATE), and Al-activated citrate exudation. {ECO:0000269|PubMed:17535918, ECO:0000269|PubMed:18826429, ECO:0000269|PubMed:19321711, ECO:0000269|PubMed:23935008}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
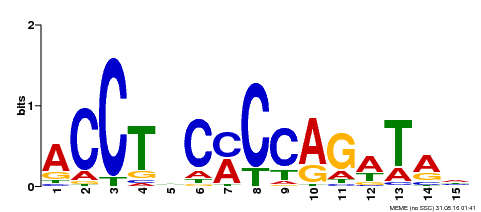
| Motif ID | Method | Source | Motif file |
| MP00196 | ampDAP | Transfer from AT1G34370 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KZV48886.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By shock H(+) and Al(3+) treatments. {ECO:0000269|PubMed:17535918}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011097108.1 | 0.0 | LOW QUALITY PROTEIN: protein SENSITIVE TO PROTON RHIZOTOXICITY 1 | ||||
| Swissprot | Q9C8N5 | 1e-179 | STOP1_ARATH; Protein SENSITIVE TO PROTON RHIZOTOXICITY 1 | ||||
| TrEMBL | A0A2Z7CVN1 | 0.0 | A0A2Z7CVN1_9LAMI; Protein SENSITIVE TO PROTON RHIZOTOXICITY 1-like | ||||
| STRING | XP_008237278.1 | 0.0 | (Prunus mume) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4754 | 24 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G34370.2 | 1e-158 | C2H2 family protein | ||||



