 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KZV39952.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Gesneriaceae; Didymocarpoideae; Trichosporeae; Loxocarpinae; Dorcoceras
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 545aa MW: 60296.1 Da PI: 5.9763 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 58.1 | 2.1e-18 | 39 | 86 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WTt Ed +l+d+vk++G g+W+++ ++ g+ R++k+c++rw ++l
KZV39952.1 39 KGPWTTAEDAILIDYVKKHGEGNWNSVQKHTGLSRCGKSCRLRWANHL 86
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 51.4 | 2.5e-16 | 92 | 135 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T+eE+ l++++++++G++ W++ a +++ gRt++++k++w++
KZV39952.1 92 KGAFTPEEERLIIELHAKMGNK-WARMAVHLP-GRTDNEIKNYWNT 135
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 18.018 | 34 | 86 | IPR017930 | Myb domain |
| SMART | SM00717 | 7.1E-16 | 38 | 88 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 9.36E-31 | 38 | 133 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 1.9E-16 | 39 | 86 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.4E-25 | 40 | 94 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.92E-12 | 41 | 86 | No hit | No description |
| PROSITE profile | PS51294 | 25.82 | 87 | 141 | IPR017930 | Myb domain |
| SMART | SM00717 | 7.7E-17 | 91 | 139 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.2E-15 | 92 | 135 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.23E-12 | 94 | 135 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 3.9E-25 | 95 | 140 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 545 aa Download sequence Send to blast |
MGHSTGDERE NRALSCDSPL SNDGDNFLRG SDRGVILKKG PWTTAEDAIL IDYVKKHGEG 60 NWNSVQKHTG LSRCGKSCRL RWANHLRPNL KKGAFTPEEE RLIIELHAKM GNKWARMAVH 120 LPGRTDNEIK NYWNTRIKRR QRAGLPLYPA EFFQQALPEN QQSRNSPGIF GLDEGCHDSL 180 PSHGYETPDV MFDSLNMLPY RAEFPDISAS GSLVTGFGSP QLYTFLPQTV GLHKLTQEHD 240 EQMDFGCSAV NGSYPFDMAR NDRCENKMSQ SYGLYLPSDP NHTKQLWPFG VSPDSHFTSN 300 DIFSASEPLE GAEKLELPSL QYQETNSGVW GLLPFDAAPE SLDSFIQSPL SGTHPSHCSS 360 PRSSGLLEAL LYEAKFRSKS ENQSFDKATS STVSQGAMTY CSSLKAHSTE IKGVDPTSPS 420 GNSTGSNFNR CSPVDPIGTA MEEKIRASRM KSEVVNQRCI VILDEDRKEN LNHLDCSPPD 480 AVLGSCWLEQ SGSTVKQRGS ITEAIFLGDD LGSELKNMSS RTSGWGLGSC TWNNMPSAYQ 540 MSDIQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 3e-32 | 37 | 140 | 5 | 107 | B-MYB |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
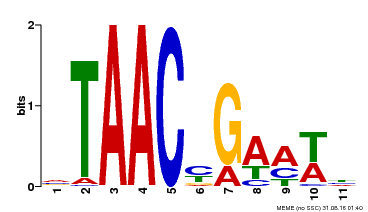
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KZV39952.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011091869.1 | 0.0 | transcription factor GAMYB | ||||
| Refseq | XP_011091870.1 | 0.0 | transcription factor GAMYB | ||||
| Refseq | XP_011091873.1 | 0.0 | transcription factor GAMYB | ||||
| Refseq | XP_020552724.1 | 0.0 | transcription factor GAMYB | ||||
| TrEMBL | A0A2Z7BZ70 | 0.0 | A0A2Z7BZ70_9LAMI; Transcription factor GAMYB-like | ||||
| STRING | Migut.N01870.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA14059 | 7 | 13 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 1e-114 | myb domain protein 33 | ||||



