 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KZV38237.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Gesneriaceae; Didymocarpoideae; Trichosporeae; Loxocarpinae; Dorcoceras
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 325aa MW: 36828.5 Da PI: 8.4526 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 62 | 9.2e-20 | 57 | 109 | 4 | 56 |
-SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 4 RttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
++++++eq Le+ F +++++ e++ +LA++lgL+ rqV vWFqNrRa++k
KZV38237.1 57 KRKLSEEQMSMLERSFGSEHKLESERKDRLAAELGLDPRQVAVWFQNRRARWK 109
45899***********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 117.5 | 7.3e-38 | 56 | 147 | 2 | 93 |
HD-ZIP_I/II 2 kkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
+kr+ls+eq+++LE+sF +e+kLe+erK +la eLgl+prqvavWFqnrRAR+k+k+lE++y++Lk+++d+ + e++rLe++ +L+e+l+e
KZV38237.1 56 RKRKLSEEQMSMLERSFGSEHKLESERKDRLAAELGLDPRQVAVWFQNRRARWKSKKLEEEYSRLKSEHDSTVVEKCRLETQLLKLKEQLRE 147
8****************************************************************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 16.86 | 51 | 111 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 4.1E-18 | 54 | 115 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.44E-16 | 56 | 112 | No hit | No description |
| Pfam | PF00046 | 4.2E-17 | 57 | 109 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 7.7E-18 | 57 | 120 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 6.0E-20 | 58 | 118 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 1.8E-5 | 82 | 91 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 86 | 109 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.8E-5 | 91 | 107 | IPR000047 | Helix-turn-helix motif |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 325 aa Download sequence Send to blast |
MNPETDDPMA LISEYYPSEI YSQFEPQQEA AAPNPKPRRR RKKNRGDEGG VDGVMRKRKL 60 SEEQMSMLER SFGSEHKLES ERKDRLAAEL GLDPRQVAVW FQNRRARWKS KKLEEEYSRL 120 KSEHDSTVVE KCRLETQLLK LKEQLREAEK EIRRLSECGD GFSSNSPSSS FSMDVMNKPP 180 FMGEFGMEGL ENVIYNGSDC NNYGLGLGIT DSACKNQLVV VSVQYGPFNT HIPIRSTTIG 240 KSRVAKDPIA MHTSWRSNSD IASVTRQTHL PKTYPVLWPE IYHTIVSSDS IGYPCMRASG 300 ESSTTKHRLL HASGPHPIPP PNDPK |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 40 | 58 | RKKNRGDEGGVDGVMRKRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00473 | DAP | Transfer from AT4G36740 | Download |
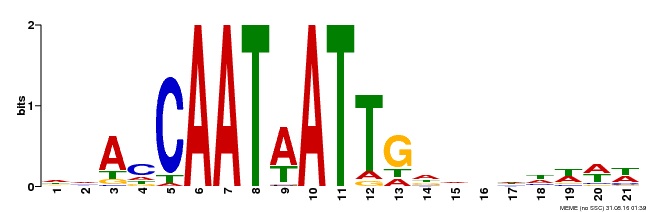
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KZV38237.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By abscisic acid (ABA) and by salt stress. {ECO:0000269|PubMed:16055682}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011099589.2 | 2e-88 | homeobox-leucine zipper protein ATHB-40 | ||||
| Swissprot | O23208 | 7e-62 | ATB40_ARATH; Homeobox-leucine zipper protein ATHB-40 | ||||
| TrEMBL | A0A2Z7C140 | 0.0 | A0A2Z7C140_9LAMI; Homeobox-leucine zipper protein ATHB-40 | ||||
| STRING | Solyc02g085630.2.1 | 4e-76 | (Solanum lycopersicum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G36740.1 | 9e-48 | homeobox protein 40 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



