 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KZV25835.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Gesneriaceae; Didymocarpoideae; Trichosporeae; Loxocarpinae; Dorcoceras
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 312aa MW: 35395.6 Da PI: 4.9531 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 27.7 | 4.6e-09 | 251 | 283 | 23 | 55 |
SS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 23 rypsaeereeLAkklgLterqVkvWFqNrRake 55
+yp++ee+ +L++ +gL+++q+ +WF N+R ++
KZV25835.1 251 PYPTEEEKNRLSEMTGLDQKQINNWFINQRKRH 283
8*****************************885 PP
| |||||||
| 2 | ELK | 39 | 1.7e-13 | 203 | 224 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELK++L+rKYsgyL+sL++EF+
KZV25835.1 203 ELKQMLMRKYSGYLSSLRKEFL 224
9********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 5.0E-20 | 50 | 94 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 3.2E-20 | 52 | 92 | IPR005540 | KNOX1 |
| SMART | SM01256 | 9.5E-27 | 98 | 149 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 3.7E-25 | 101 | 148 | IPR005541 | KNOX2 |
| SMART | SM01188 | 1500 | 107 | 127 | IPR005539 | ELK domain |
| SMART | SM01188 | 2.8E-7 | 203 | 224 | IPR005539 | ELK domain |
| Pfam | PF03789 | 2.3E-10 | 203 | 224 | IPR005539 | ELK domain |
| PROSITE profile | PS51213 | 11.366 | 203 | 223 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.681 | 223 | 286 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.03E-18 | 225 | 295 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 1.3E-11 | 225 | 290 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 2.5E-25 | 228 | 288 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.20E-12 | 239 | 286 | No hit | No description |
| Pfam | PF05920 | 7.9E-17 | 243 | 282 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 261 | 284 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 312 aa Download sequence Send to blast |
MDELYGFHCS VDVGFNGDGN SIGVEQMVLT AQVAEIHGGL TSGITSEMSD IIKAQIAYHP 60 LYPHLLSAYV ECRKVGAPPE MVSLLEEISR ENYPLSSCSS EIGADPELDE FMRSYCEVLH 120 KYKEELSRPF DEATSFLSSI ESQLNDLCKE TLPSVTNNSS LTPPSKYHSA DDVGGRSDED 180 ASCGELEASE SLESCFNQQG DEELKQMLMR KYSGYLSSLR KEFLKKRKKG KLPKNARTAL 240 LHWWNTHYRW PYPTEEEKNR LSEMTGLDQK QINNWFINQR KRHWRPLEDM RFAVMEGVSG 300 GVNHNYNVID GS |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 218 | 227 | LRKEFLKKRK |
| 2 | 224 | 228 | KKRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may be involved in shoot formation during early embryogenesis. {ECO:0000269|PubMed:10488233}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00670 | PBM | Transfer from GRMZM2G087741 | Download |
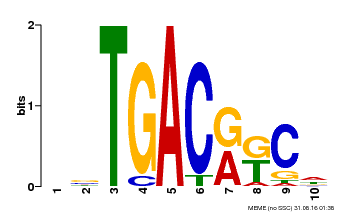
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KZV25835.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011087493.1 | 1e-152 | homeobox protein knotted-1-like 1 | ||||
| Swissprot | Q9FP29 | 5e-97 | KNOS1_ORYSJ; Homeobox protein knotted-1-like 1 | ||||
| TrEMBL | A0A2Z7AVG3 | 0.0 | A0A2Z7AVG3_9LAMI; Homeobox protein knotted-1-like 1 | ||||
| STRING | VIT_14s0060g01180.t01 | 1e-124 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA753 | 24 | 80 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G23380.1 | 7e-76 | KNOTTED1-like homeobox gene 6 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



