 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KZV25739.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Gesneriaceae; Didymocarpoideae; Trichosporeae; Loxocarpinae; Dorcoceras
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 299aa MW: 33333.4 Da PI: 8.2228 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 44 | 5e-14 | 138 | 182 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT++E++l++ + k++G+g+W+ I+r + +Rt+ q+ s+ qky
KZV25739.1 138 PWTEDEHKLFLMGLKKYGKGDWRNISRNFVITRTPTQVASHAQKY 182
8*****************************99************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 7.817 | 24 | 77 | IPR017884 | SANT domain |
| SMART | SM00717 | 6.1E-8 | 25 | 77 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 7.18E-11 | 28 | 82 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 6.05E-7 | 29 | 74 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 2.9E-12 | 129 | 187 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 20.138 | 131 | 187 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.01E-16 | 133 | 186 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.8E-17 | 134 | 186 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.3E-12 | 135 | 185 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 6.66E-11 | 138 | 183 | No hit | No description |
| Pfam | PF00249 | 1.8E-12 | 138 | 182 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 299 aa Download sequence Send to blast |
MKWGMEILSP SSCFSNSSLF LGESRATGWT QAENKAFENA LAMFDENTPD RWRRVAEMVP 60 GKTVGDVMRQ YKELEDDVSS IEAGLIPVPG YTTSLPFTLE WGGSHVYDGF NQSCHVSGGR 120 KSGSLVRSSD QERKKGVPWT EDEHKLFLMG LKKYGKGDWR NISRNFVITR TPTQVASHAQ 180 KYFIRQLSGG KDKRRASIHD ITTVTIGDNQ MLPPDDKKQP SPEFPNSASP MHELPLQWNQ 240 TNRDTIGGFN PGGILPYGAA SHGLKALGQN LSRNAAYESY LGSQNLIFQM QSELHYPNA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 1e-15 | 21 | 94 | 1 | 76 | RADIALIS |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the dorsovental asymmetry of flowers. Promotes ventral identity. {ECO:0000269|PubMed:11937495, ECO:0000269|PubMed:9118809}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00568 | DAP | Transfer from AT5G58900 | Download |
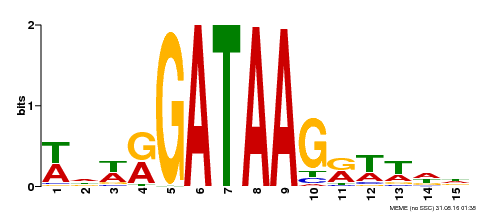
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KZV25739.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011080854.1 | 1e-147 | transcription factor DIVARICATA isoform X1 | ||||
| Refseq | XP_020550520.1 | 1e-147 | transcription factor DIVARICATA isoform X2 | ||||
| Swissprot | Q8S9H7 | 1e-143 | DIV_ANTMA; Transcription factor DIVARICATA | ||||
| TrEMBL | A0A2Z7B267 | 0.0 | A0A2Z7B267_9LAMI; Uncharacterized protein | ||||
| STRING | Gorai.009G093700.1 | 1e-133 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA458 | 24 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G58900.1 | 1e-96 | Homeodomain-like transcriptional regulator | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



