 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KZV16610.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Gesneriaceae; Didymocarpoideae; Trichosporeae; Loxocarpinae; Dorcoceras
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 839aa MW: 96210.6 Da PI: 6.9698 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 83.2 | 4.2e-26 | 85 | 187 | 2 | 91 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk.............tekerr....traetrtgCkaklkvkkekdgkwevtklel 83
fY+eY+++vGF++ +++s++sk+++e+++++f Cs++g+++e +k+ + +ra +t+Cka+++vk+++dgkw ++++e+
KZV16610.1 85 FYQEYSRSVGFNTAIQNSRRSKTSREFIDAKFACSRYGTKREYEKSlnrprsrqgskqdP----EiatgRRACAKTDCKASMHVKRRQDGKWIIHRFEK 179
9*****************************************999999998876655433....1455599**************************** PP
FAR1 84 eHnHelap 91
eHnHel p
KZV16610.1 180 EHNHELLP 187
*****975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 6.4E-24 | 85 | 187 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 3.2E-23 | 285 | 376 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.841 | 565 | 601 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.2E-6 | 576 | 603 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 839 aa Download sequence Send to blast |
MDIDLRLPSG DKEEHNGIVN MLDGDEKPLN LDGVSGSMVD VEEKLHIEDG EDINSPLHDV 60 DFKDLTILEP LPGMEFGSHG DAYAFYQEYS RSVGFNTAIQ NSRRSKTSRE FIDAKFACSR 120 YGTKREYEKS LNRPRSRQGS KQDPEIATGR RACAKTDCKA SMHVKRRQDG KWIIHRFEKE 180 HNHELLPAQA VSEQTRRMYA AMARQFAEYK SAVGLKHDSR SSFEKSRNMT MDAEEVNIML 240 EFFIQMQCQN SNFFYAVDVG EDQRLKSMLW IDAKSRHDYI YFSDVLSFDT SYVRSKYKMP 300 LILFIGVNQY YQFMLLGCAL ISDETAATYT WVMQTWLKAM GGQVPKIVIT DQDKVLKSVV 360 LEIFTSTLHV FCLWNIIGKV SETLNHVIKQ NGDFTSKFEK CIYRSGTDEE FEKRWHKLVD 420 RFGLKENELF QSLFEDREKW ATSFLKDGTL AGMSAVQRSE SVNSFFDKYV HKKTTVLEFV 480 KQYESILQDR YEEEAKAYSD TWNKQPTLKS PSPFEKHVAG VYTHAVFKKF QVEVLGAVAC 540 APKKEEKVDA TIKYKVQDFE RNQEFTVTLN ETKSEVSCIC HLFEFKGFLC RHAMIVLQIC 600 GISAIPSQYI LKRWTKDARS RYSSGDAYEL VQSRVQRYND LYHRAVKLGE EASLSQESYN 660 MTVRAIDGAF ENCLNVNNSD KNLVETGASV SPGLLCIEED FQGSKTNKRK NATKKRKVNV 720 EHDIMTVGTS ESLQQLEKLS SRQVNLDGFF GSQQVVQGMV QLNLMAPSRD NYYGSQQTIQ 780 GLGQLSSIAP AHDGYYGTHP TMHGLGQMDF FRTPNFNYGL REEQNVRSAQ LHDDAQRHA |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
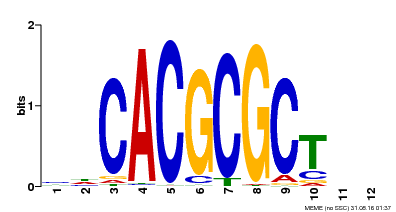
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KZV16610.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011100397.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 isoform X2 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A2Z7AC78 | 0.0 | A0A2Z7AC78_9LAMI; Protein FAR-RED ELONGATED HYPOCOTYL 3-like | ||||
| STRING | Migut.M00094.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA496 | 21 | 103 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||



