 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Dca43861.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Caryophyllaceae; Caryophylleae; Dianthus
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 1034aa MW: 112635 Da PI: 6.1925 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 93 | 2.2e-29 | 655 | 706 | 1 | 52 |
RWP-RK 1 aekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
aek+isle+l++yF++++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rki+++
Dca43861.1 655 AEKQISLEVLQQYFAGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKINKV 706
589***********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.723 | 645 | 726 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 4.0E-26 | 659 | 706 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 5.56E-22 | 932 | 1024 | No hit | No description |
| Pfam | PF00564 | 1.2E-18 | 939 | 1020 | IPR000270 | PB1 domain |
| PROSITE profile | PS51745 | 23.8 | 939 | 1021 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 1.4E-24 | 939 | 1020 | No hit | No description |
| SMART | SM00666 | 1.5E-23 | 939 | 1021 | IPR000270 | PB1 domain |
| CDD | cd06407 | 3.79E-35 | 940 | 1020 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1034 aa Download sequence Send to blast |
MSQTPPATTN TNTHPDSDTN NNNNNNNNNN NNKSCFGVGV GVGVGPTATS SRGENYYYSS 60 GSGIHHNNNN NNNNNNNNIY VNLEASSTAL MDLDFGDLDA SWSFDQITAA GGAGGGAAAT 120 ITAAPVLISN SDQPCSPLWA FSDAPHFPNS FCFSPCHPDP ISHTATHTCG VNAPLHDDDD 180 HNNNNKIPLP FTALSPPDNL DVSFLIKERM TRALRYFKEL TEQHQILAQV WAPVRSGNRY 240 VLTTSGQPFV LGPHSNGLNQ YRTVSLMYMF AVDGDSDVTL GLPGRVFNQK LPEWTPNVQY 300 YSSKEYPRVD HAKHYNVKGT LALPVFEPSG QSCVGVLELI MTSQKINYAP EVDKVCKALE 360 AVNLRSSEIL DHPNIQICNE GRQNALAEIL EVLTEVCETY KLPLAQTWIP CKHRSVLAYG 420 GGLKKSCSSF DGSCMGRVCM STTDVAYYVV DAQILGFREA CAEHHLQRGQ GVAGRAFSSH 480 GLSFCRDISQ FCKFDYPLVH YARMFRFRGS FAICLRSKHT GDDDYVLEFF LPSSITDSAG 540 QLDLLDSILA TMKEDLQNLM FASGNTLEEE RKHMEIIEVS TDYNHDAQIG SNLSIIPTSM 600 AMAMNGALSL AKDSQKQLTA EAGAMNIVAS RSHNPISPTG KKEAKKTSER KRGKAEKQIS 660 LEVLQQYFAG SLKDAAKSLG VCPTTMKRIC RQHGISRWPS RKINKVNRSL TKLKRVIESV 720 QGADGTFSLT SLAQPTMPVP VGSISWPAGV NGTCHQLSPT SQPTDVPDER SEMPSYEPRT 780 NGQTCGRASS REDRVNEQNG FTSKGGSGST GTSTSHGSCH GSPGTESNPP PNGNFVQSFC 840 EQVGNSSGLM FRQNGAYSIP DALFIAHSQD TFGGMLLEDA GSSKDLRNIC SPGVEANADE 900 SPAPEAGWIN PSCPSPSSRV GTSVFGMTSR FSVRQDMKTV TIKATFREDI IRFRFSLASN 960 IVELREEVSK RLKLEVGTFE IKYLDDDHEW VLIACDSDLQ ECLDVSRSSG SDMIRLLIQD 1020 IMGNLGSSCE SSGE |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
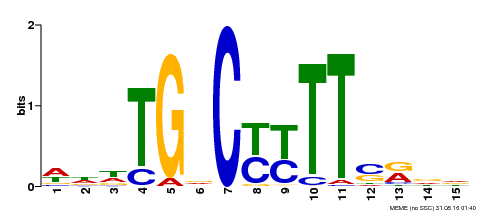
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010695590.1 | 0.0 | PREDICTED: protein NLP7 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | A0A0J8BC07 | 0.0 | A0A0J8BC07_BETVU; Uncharacterized protein | ||||
| STRING | XP_010695590.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||



