 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | DCAR_018677 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; campanulids; Apiales; Apiineae; Apiaceae; Apioideae; Scandiceae; Daucinae; Daucus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 352aa MW: 39680.9 Da PI: 6.4198 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 100.3 | 1.3e-31 | 66 | 119 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWt++LH rFv+ave+LGG+e+AtPk +l+lm+vkgL ++hvkSHLQ+YR+
DCAR_018677 66 PRLRWTHDLHLRFVQAVERLGGQERATPKLVLQLMNVKGLNIAHVKSHLQMYRS 119
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 13.164 | 62 | 122 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 8.5E-30 | 62 | 120 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 5.91E-15 | 63 | 120 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 4.4E-23 | 66 | 121 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 3.1E-8 | 67 | 118 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 352 aa Download sequence Send to blast |
MDGSSRSECY KASPSNEEYE EDEEDVSAEN NSKATDGGAG GSSSNSTIEE NENKSSVRPY 60 VRSKTPRLRW THDLHLRFVQ AVERLGGQER ATPKLVLQLM NVKGLNIAHV KSHLQMYRSK 120 KIDDPNQAIA DNRSVENYGD RNIYNLSQLP MLQSYNQSYN LSTFRYGDAL WNAGHHEKWM 180 QNVYNMSGRN GAVDYKARPR FYPSVTEKIS GRNSINWENN INASSSSQLS TWKVHENKGE 240 LLGPLNDQSP VKFMEFKSSN RQQEKVEPLD LEIASLTPAK RKASACELDL NLTLAVKSTN 300 DNSGRNLKDD EDEYLSLSLF SPSSSKKLER LKMVDKEEIN AERASTLDLT I* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 2e-16 | 67 | 121 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 2e-16 | 67 | 121 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 2e-16 | 67 | 121 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 2e-16 | 67 | 121 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 2e-16 | 67 | 121 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 2e-16 | 67 | 121 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 2e-16 | 67 | 121 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 2e-16 | 67 | 121 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 2e-16 | 67 | 121 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 2e-16 | 67 | 121 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 2e-16 | 67 | 121 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 2e-16 | 67 | 121 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00297 | DAP | Transfer from AT2G38300 | Download |
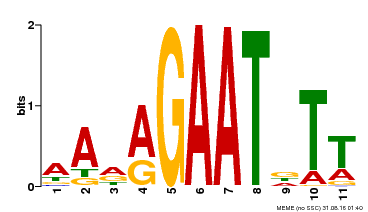
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017251636.1 | 0.0 | PREDICTED: putative Myb family transcription factor At1g14600 | ||||
| TrEMBL | A0A164Z6V7 | 0.0 | A0A164Z6V7_DAUCS; Uncharacterized protein | ||||
| STRING | EOY07117 | 2e-89 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4382 | 21 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38300.1 | 1e-53 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | DCAR_018677 |



