 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | orange1.1g048771m | ||||||||
| Common Name | CISIN_1g048771mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 815aa MW: 89651.8 Da PI: 6.6639 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58.5 | 1.1e-18 | 17 | 75 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++++++ps +r++L +++ +++ +q+kvWFqNrR +ek+
orange1.1g048771m 17 KYVRYTPEQVEALERLYHECPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 75
5679*****************************************************97 PP
| |||||||
| 2 | START | 34.5 | 5e-12 | 161 | 227 | 2 | 69 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCG CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddk 69
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s+++sg a+ra+g+v +++ v+e+l+d+
orange1.1g048771m 161 IAEETLTEFLSKATGTAVEWVQMPGMKPGPDSVGIVAISHGCSGVAARACGLVGLEPT-RVAEILKDR 227
789*******************************************************.778877775 PP
| |||||||
| 3 | START | 104.1 | 2.3e-33 | 234 | 345 | 98 | 205 |
XTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHH CS
START 98 lqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslv 185
l+al++l+p Rdf+ +Ry+ l++g++v++++S+ + q+ p+ +++vRae+lpSg+li+p+++g+s +++v+h+dl+ ++++++lr+l+
orange1.1g048771m 234 LYALTTLAPaRDFWLLRYTSVLEDGSLVVCERSLKNIQNGPTmppVQHFVRAEMLPSGYLIRPCEGGGSIIHIVDHMDLEPWSVPEVLRPLY 325
6899********************************99999988899********************************************* PP
HHHHHHHHHHHHHHTXXXXX CS
START 186 ksglaegaktwvatlqrqce 205
+s+++ ++kt++a+l+++++
orange1.1g048771m 326 ESSTVLAQKTTMAALRQLRQ 345
***************99875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.613 | 12 | 76 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.2E-15 | 14 | 80 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 3.46E-17 | 16 | 80 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 8.87E-17 | 17 | 77 | No hit | No description |
| Pfam | PF00046 | 2.8E-16 | 18 | 75 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-18 | 19 | 75 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 1.11E-6 | 69 | 108 | No hit | No description |
| PROSITE profile | PS50848 | 18.796 | 158 | 355 | IPR002913 | START domain |
| SMART | SM00234 | 8.2E-26 | 160 | 346 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.65E-30 | 160 | 348 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 2.1E-16 | 161 | 343 | IPR023393 | START-like domain |
| Pfam | PF01852 | 2.0E-9 | 161 | 228 | IPR002913 | START domain |
| Pfam | PF01852 | 3.1E-30 | 234 | 345 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 4.07E-5 | 373 | 465 | No hit | No description |
| SuperFamily | SSF55961 | 4.07E-5 | 497 | 572 | No hit | No description |
| Pfam | PF08670 | 6.9E-52 | 671 | 813 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 815 aa Download sequence Send to blast |
MAMSCKDGKT GSLDNGKYVR YTPEQVEALE RLYHECPKPS SIRRQQLIRE CPILSNIEPK 60 QIKVWFQNRR CREKQRKEAS RLQAVNRKLT AMNKLLMEEN DRLQKQVSQL VYENGYFRQH 120 TQSTTLATKD TSCESVVTSG QHHLTPQHPP RDASPAGLLS IAEETLTEFL SKATGTAVEW 180 VQMPGMKPGP DSVGIVAISH GCSGVAARAC GLVGLEPTRV AEILKDRPRG SAILYALTTL 240 APARDFWLLR YTSVLEDGSL VVCERSLKNI QNGPTMPPVQ HFVRAEMLPS GYLIRPCEGG 300 GSIIHIVDHM DLEPWSVPEV LRPLYESSTV LAQKTTMAAL RQLRQMAQEV TQSSVNGWGR 360 RPAALRALSQ RLSRGFNEAV NGFTDEGWTV MGNDGMDDVT VLVNSSPDKL MGLNLSFANG 420 FPAVSNAVLC AKASMLLQNV PPAILLRFLR EHRSEWADNN IDVYSAAAIK VGPCSLPGSR 480 VGTFGSQVIL PLAHTIEHEE FMEVIKLEGV GHSPEDAIMP RDMFLLQLCS GMDENAVGTC 540 AELIFAPIDA SFADDAPLLP SGFRIIPLDS GKETSSPNRT LDLASALEIG PAGNRATNNY 600 STNSTCMRSV MTIAFEFAFE SHMQEHVATM ARQYVRSIIS SVQRVALALS PSNISSQAGL 660 RTPLGTPEAL TLARWICHSY RCYLGVDLLK SSSEGSESIL KNLWHHTDAV MCCSLKALPV 720 FTFANQAGLD MLETTLVALQ DITLEKIFDD HGRKALFAEF PQIMQQGFAC LQGGICLSSM 780 GRPVSYERAV AWKVLNEEET AHCICFMFIN WSFV* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Csi.1079 | 0.0 | callus| fruit| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed the developing vascular elements and the adaxial portion of cotyledons. Expressed in developing ovules, stamens and carpels. Expressed in procambium and shoot meristem. {ECO:0000269|PubMed:14701930, ECO:0000269|PubMed:15705957}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
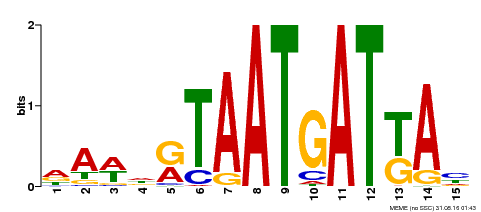
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006426893.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Refseq | XP_006465680.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Refseq | XP_015386901.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A067ED21 | 0.0 | A0A067ED21_CITSI; Uncharacterized protein | ||||
| STRING | XP_006465679.1 | 0.0 | (Citrus sinensis) | ||||
| STRING | XP_006426892.1 | 0.0 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2022 | 26 | 78 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | orange1.1g048771m |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



