 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | orange1.1g025053m | ||||||||
| Common Name | CISIN_1g021635mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 259aa MW: 29146.3 Da PI: 10.0524 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 109.1 | 5.2e-34 | 20 | 160 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkk......lelee...vikev.diykvePwdLpkkvkaeekewyfFskrdkkyatgkrknratk... 79
lp+G++F+Ptd+el+ e+L++kv+++ ++ e +++ +i++++P++Lp v+++ +fF++ +k+y+tg+rk+r+++
orange1.1g025053m 20 LPAGVKFDPTDQELL-EHLEAKVRADArklhplID--EfipTLEGEnGICYTHPEKLP-GVSKDGLIRHFFHRPSKAYTTGTRKRRKVHtde 107
799************.99*****999956643322..22222333336**********.555555677******************744456 PP
NAM 80 ...sgyWkatgkdkevlskkgelvglkktLvfy..kgrapkgektdWvmheyrl 128
+ +W++tgk+++v+ +g+++g kk+Lv+y g+++k+ekt+Wvmh+y+l
orange1.1g025053m 108 qggETRWHKTGKTRPVFI-GGKVKGYKKILVLYtnYGKQKKPEKTNWVMHQYHL 160
555778************.9*************6658999************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 1.57E-38 | 14 | 179 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 33.694 | 20 | 179 | IPR003441 | NAC domain |
| Pfam | PF02365 | 2.8E-16 | 21 | 160 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:2000652 | Biological Process | regulation of secondary cell wall biogenesis | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 259 aa Download sequence Send to blast |
MVIMIRWPFF KARIHDLPGL PAGVKFDPTD QELLEHLEAK VRADARKLHP LIDEFIPTLE 60 GENGICYTHP EKLPGVSKDG LIRHFFHRPS KAYTTGTRKR RKVHTDEQGG ETRWHKTGKT 120 RPVFIGGKVK GYKKILVLYT NYGKQKKPEK TNWVMHQYHL GGNEEEKDGE LVVSKVFYQT 180 QPRQCGNSVI KDSSSLPPKF NINKGQSGLD GLKNNPGVVD YYNPPFIAFD QGGGQNRASP 240 PQALIPPHFA AHDTSFIP* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 8e-12 | 20 | 160 | 15 | 140 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in protoxylem and elongating interfascicular fiber cells of elongating internodes, developing metaxylem cells and interfascicular fibers of non-elongating internodes and developing secondary xylem of roots. {ECO:0000269|PubMed:18952777}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that plays a regulatory role in the development of secondary cell wall fibers (PubMed:18952777,PubMed:22133261). Involved in the regulation of cellulose and hemicellulose biosynthetic genes as well as of genes involved in lignin polymerization and signaling (PubMed:22133261). Is not a direct target of SND1 (PubMed:18952777). {ECO:0000269|PubMed:18952777, ECO:0000269|PubMed:22133261}. | |||||
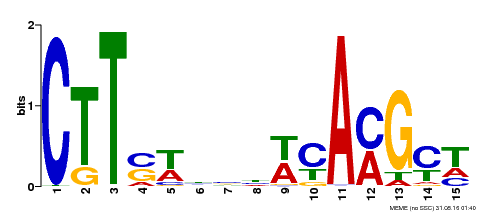
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00459 | DAP | Transfer from AT4G28500 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006469079.1 | 0.0 | NAC domain-containing protein 73 | ||||
| Swissprot | O49459 | 1e-113 | NAC73_ARATH; NAC domain-containing protein 73 | ||||
| TrEMBL | A0A067E6S0 | 0.0 | A0A067E6S0_CITSI; Uncharacterized protein | ||||
| TrEMBL | A0A2H5PT91 | 0.0 | A0A2H5PT91_CITUN; Uncharacterized protein | ||||
| STRING | XP_006469079.1 | 0.0 | (Citrus sinensis) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G28500.1 | 1e-115 | NAC domain containing protein 73 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | orange1.1g025053m |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



