 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | orange1.1g024464m | ||||||||
| Common Name | CISIN_1g024344mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 268aa MW: 29530.7 Da PI: 7.8853 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 33.3 | 8.8e-11 | 152 | 199 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+h+ +Er RR++i +++ L++l+P + +k Ka +L + ++YI+sLq
orange1.1g024464m 152 SHSLAERARREKISERMKILQDLVPGC----NKVIGKALVLDEIINYIQSLQ 199
8*************************9....677*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00083 | 1.58E-9 | 146 | 201 | No hit | No description |
| SuperFamily | SSF47459 | 9.29E-16 | 146 | 210 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 15.828 | 148 | 198 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 4.3E-16 | 148 | 210 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.8E-8 | 152 | 199 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.7E-8 | 154 | 204 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048446 | Biological Process | petal morphogenesis | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 268 aa Download sequence Send to blast |
MDQGSYNFAE IWQFPVPGSG SMSESGGGLG QKGAHFGQHL SQFGTNREVS GDDPVNLEHK 60 MAHGNGVRKR RDVEDESAKH VSTSSGNGNG NRVNDSDGKR IKTSGSRDDN HHSKAEAEPS 120 SVKPAEQNSQ PPEPPKQDYI HVRARRGQAT DSHSLAERAR REKISERMKI LQDLVPGCNK 180 VIGKALVLDE IINYIQSLQR QFLSMKLEAV NTRMNPGIEV FPPKDFTQQT FDTAGMPFVS 240 QATREYSRGT SPDWLHMQIG GGFERMT* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that may regulate jasmonate-regulated genes. {ECO:0000250|UniProtKB:Q69WS3}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00207 | DAP | Transfer from AT1G59640 | Download |
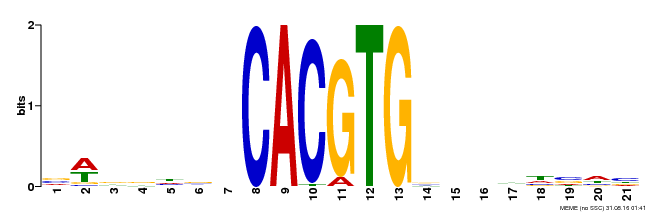
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006438535.1 | 0.0 | transcription factor BHLH089 isoform X1 | ||||
| Refseq | XP_006483289.1 | 0.0 | transcription factor BHLH089 isoform X1 | ||||
| Swissprot | Q84T08 | 7e-80 | BH089_ORYSJ; Transcription factor BHLH089 | ||||
| TrEMBL | A0A067GSM6 | 0.0 | A0A067GSM6_CITSI; Uncharacterized protein | ||||
| STRING | XP_006483289.1 | 0.0 | (Citrus sinensis) | ||||
| STRING | XP_006438535.1 | 0.0 | (Citrus clementina) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G59640.1 | 9e-91 | BIG PETAL P | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | orange1.1g024464m |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



