 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | orange1.1g019599m | ||||||||
| Common Name | CISIN_1g0195702mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 339aa MW: 38207.8 Da PI: 4.4086 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.4 | 6e-19 | 60 | 113 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ +q+++Le+ Fe ++++ e++ +LA++lgL+ rqV vWFqNrRa++k
orange1.1g019599m 60 KKRRLSVDQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWK 113
566899***********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 130.6 | 6e-42 | 59 | 151 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelk 92
ekkrrls +qvk+LE++Fe e+kLeperKv+la+eLglqprqvavWFqnrRAR+ktkqlE+dy +Lk++ydalk + ++L++++e+L +e++
orange1.1g019599m 59 EKKRRLSVDQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWKTKQLERDYGVLKANYDALKLNYDSLQHDNEALLKEIR 150
69*************************************************************************************99998 PP
HD-ZIP_I/II 93 e 93
e
orange1.1g019599m 151 E 151
6 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.82E-19 | 50 | 117 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.443 | 55 | 115 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 6.3E-18 | 58 | 119 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 3.9E-16 | 60 | 113 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 8.33E-17 | 60 | 116 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 7.1E-20 | 62 | 122 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 9.9E-6 | 86 | 95 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 90 | 113 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 9.9E-6 | 95 | 111 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 3.2E-18 | 115 | 156 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009788 | Biological Process | negative regulation of abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 339 aa Download sequence Send to blast |
MKRSLCSSDD SLGALMSICP ATDEQSPRNN QVYSREFQTM LDGLDEEGCL EESGGHVSEK 60 KRRLSVDQVK ALEKNFEVEN KLEPERKVKL AQELGLQPRQ VAVWFQNRRA RWKTKQLERD 120 YGVLKANYDA LKLNYDSLQH DNEALLKEIR ELKSKLNEEN TESNISVKQE ETIVHETDHQ 180 NKATLDRDQE SDDKQAAAVA PPTNVTAISL APAGNISDEP DQELNYDNGV LGISLFPDLK 240 DGSSDSDSSA ILNNEDNNNF HNSNNSHSPN AAISSSDGVI IQSQPHQLSI NCFEFSKPTY 300 QTQFVKMEEH NFFGDETCNF FSDEQPPSLP WYCSDQWT* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 107 | 115 | RRARWKTKQ |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Csi.717 | 0.0 | flower| fruit| seed | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Widely expressed with a lower level in siliques. {ECO:0000269|PubMed:16055682}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may function as a negative regulator of the flowering time response to photoperiod. May act to repress cell expansion during plant development. {ECO:0000269|PubMed:14623244}. | |||||
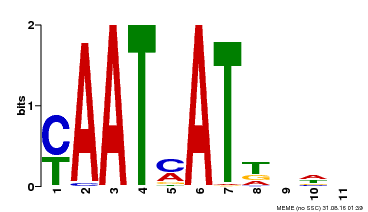
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00271 | DAP | Transfer from AT2G22430 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006426320.1 | 0.0 | homeobox-leucine zipper protein ATHB-6 | ||||
| Swissprot | Q940J1 | 4e-78 | ATB16_ARATH; Homeobox-leucine zipper protein ATHB-16 | ||||
| TrEMBL | A0A067EZZ7 | 0.0 | A0A067EZZ7_CITSI; Uncharacterized protein | ||||
| TrEMBL | V4SL15 | 0.0 | V4SL15_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006426320.1 | 0.0 | (Citrus clementina) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22430.1 | 2e-71 | homeobox protein 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | orange1.1g019599m |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



