 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | orange1.1g016585m | ||||||||
| Common Name | CISIN_1g016585mg, LOC102624045 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 388aa MW: 42910.3 Da PI: 7.4803 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.5 | 5.3e-32 | 217 | 272 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r+ W+peLH+rF++a++qLGGs+ AtPk+i+elm+v+gLt+++vkSHLQkYRl+
orange1.1g016585m 217 KQRRSWSPELHRRFLHALQQLGGSHAATPKQIRELMNVDGLTNDEVKSHLQKYRLH 272
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 13.438 | 214 | 274 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.0E-28 | 215 | 275 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.97E-18 | 215 | 275 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 9.5E-27 | 217 | 272 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 3.9E-8 | 219 | 270 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 388 aa Download sequence Send to blast |
MDYAEKMQRC HEYVEALEEE KRKIQVFQRE LPLCLELVTQ AIQACRRELS GTTTEYMHVQ 60 SECSEQTSSD GPILEEFIPI KRSSSHDDND NDNDDRHSNR PTTTNNNREK EKNSDKKKSD 120 WLRSVQLWNQ SPDPQPKEEE VVPRKATVVE VKRNGSGGAF QPFHREKCGE KKDQQQRLGK 180 VNTPVPTAAA SSTTDTAVGK GEKNDKDKDK EGQPQRKQRR SWSPELHRRF LHALQQLGGS 240 HAATPKQIRE LMNVDGLTND EVKSHLQKYR LHTRRPSPAM HNGGNPQAPQ FVVVGGIWVQ 300 APDYAAVAAK APGEAGSIAT SNGIYAPVAA PPPAVPQSSA SVVQISQQTQ SEHLQSEERG 360 SHSDGGHHAN SPATSSSTHT TTTSPVF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 2e-12 | 217 | 270 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 2e-12 | 217 | 270 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 2e-12 | 217 | 270 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 2e-12 | 217 | 270 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 2e-12 | 217 | 270 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 2e-12 | 217 | 270 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 2e-12 | 217 | 270 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 2e-12 | 217 | 270 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 2e-12 | 217 | 270 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 2e-12 | 217 | 270 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 2e-12 | 217 | 270 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 2e-12 | 217 | 270 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in phosphate signaling in roots. {ECO:0000250|UniProtKB:Q9FX67}. | |||||
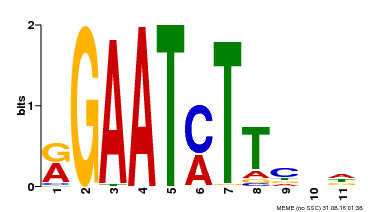
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006484416.1 | 0.0 | transcription factor HHO3 | ||||
| Swissprot | Q9FPE8 | 1e-115 | HHO3_ARATH; Transcription factor HHO3 | ||||
| TrEMBL | A0A067G303 | 0.0 | A0A067G303_CITSI; Uncharacterized protein | ||||
| TrEMBL | A0A2H5P3T6 | 0.0 | A0A2H5P3T6_CITUN; Uncharacterized protein | ||||
| STRING | XP_006484416.1 | 0.0 | (Citrus sinensis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3491 | 28 | 63 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25550.1 | 9e-95 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | orange1.1g016585m |
| Entrez Gene | 102624045 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



