 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | orange1.1g010053m | ||||||||
| Common Name | CISIN_1g010053mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 520aa MW: 58204.6 Da PI: 6.7995 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 41.9 | 1.8e-13 | 342 | 387 | 4 | 54 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksL 54
+h e+Er+RR+r+N++f Lr+++P+ K++Ka+ L Av YIk+L
orange1.1g010053m 342 NHVEAERQRRERLNHRFYALRSVVPNV-----SKMDKASLLADAVAYIKEL 387
799***********************6.....5****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 2.5E-49 | 24 | 210 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| SuperFamily | SSF47459 | 2.88E-18 | 337 | 401 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 17.606 | 338 | 387 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 6.66E-15 | 341 | 391 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 1.5E-17 | 342 | 403 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 5.9E-11 | 342 | 387 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 6.9E-17 | 344 | 393 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0042538 | Biological Process | hyperosmotic salinity response | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005509 | Molecular Function | calcium ion binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 520 aa Download sequence Send to blast |
MEEIVSSSSS SYPMPFCQET SPTLQQRLQF IVQNRPEWWV YSIFWQPLKD VNGRLVLSWG 60 DGYFRGSKDF ATRAAAGKQG AGNEPKFGFF LERKKVSKEV QVHFGEDMDL DRMVDGDVTD 120 GEWYYTVSVT RSFAIGDGSV LGRVFSSGDY VWLTGDHELQ LYECERVKEA RMHGIQTLVC 180 VSTACGVVEL GSSDLIKEDW SLVQLAKSLF GPVIATMLTK QVNLNSESQL QLPNPTTRNN 240 NNTNNVAPLL DIGMFSGAGA PHHHHHHHQK EWSLEENSKQ QTREVSGDVI KKEQLAAGFG 300 RSSSDSGPSD SDGHFVSGFT DINVTSKKRG RKPTSGRESP LNHVEAERQR RERLNHRFYA 360 LRSVVPNVSK MDKASLLADA VAYIKELRAK VDELEAKLRE QARKSKVVYN VYDNNQSTGS 420 TIMMPTSSST THHLGININI MDVDVKIVGS EAMIRVQCPD INYPAAKLMD VLRDLEFHVH 480 HASVSSVRET MLQDVVVRIP EGLISEEVIR SAIFQRMQN* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4rqw_A | 1e-26 | 18 | 210 | 3 | 192 | Transcription factor MYC3 |
| 4rqw_B | 1e-26 | 18 | 210 | 3 | 192 | Transcription factor MYC3 |
| 4rs9_A | 1e-26 | 18 | 210 | 3 | 192 | Transcription factor MYC3 |
| 4yz6_A | 1e-26 | 18 | 210 | 3 | 192 | Transcription factor MYC3 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 347 | 352 | RQRRER |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Csi.7566 | 0.0 | flower| fruit | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00547 | DAP | Transfer from AT5G46830 | Download |
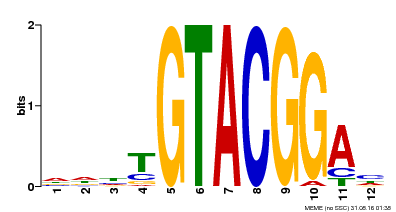
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006476285.1 | 0.0 | transcription factor MYC2-like | ||||
| TrEMBL | A0A067GDM6 | 0.0 | A0A067GDM6_CITSI; Uncharacterized protein | ||||
| STRING | XP_006476285.1 | 0.0 | (Citrus sinensis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3039 | 21 | 63 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G46830.1 | 5e-59 | NACL-inducible gene 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | orange1.1g010053m |



