 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | orange1.1g007393m | ||||||||
| Common Name | CISIN_1g007393mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 606aa MW: 67617.7 Da PI: 4.9388 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 169.1 | 1.5e-52 | 9 | 136 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp..kkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdk 90
lp GfrF+Ptdeelv++yL+ k++g++ e+ +vi+e+d++k ePwdLp + +k++++ew+fF+++d+ky++g+r nrat++gyWkatgkd+
orange1.1g007393m 9 LPLGFRFRPTDEELVDFYLRMKINGHNDEV-SVISEIDVCKREPWDLPdlSVIKTKDREWFFFCPQDRKYPNGHRLNRATAAGYWKATGKDR 99
699************************999.99**************9644889999*********************************** PP
NAM 91 evlskkgelvglkktLvfykgrapkgektdWvmheyrl 128
+++s +++l+g+kktLvf+ grapkg++t+Wvmheyr+
orange1.1g007393m 100 KIKS-GNHLIGMKKTLVFHMGRAPKGKRTNWVMHEYRA 136
****.99*****************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 3.4E-61 | 4 | 159 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 56.812 | 9 | 159 | IPR003441 | NAC domain |
| Pfam | PF02365 | 5.6E-27 | 10 | 135 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009814 | Biological Process | defense response, incompatible interaction | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0070417 | Biological Process | cellular response to cold | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005886 | Cellular Component | plasma membrane | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 606 aa Download sequence Send to blast |
MAVLSLNSLP LGFRFRPTDE ELVDFYLRMK INGHNDEVSV ISEIDVCKRE PWDLPDLSVI 60 KTKDREWFFF CPQDRKYPNG HRLNRATAAG YWKATGKDRK IKSGNHLIGM KKTLVFHMGR 120 APKGKRTNWV MHEYRATTDD LDGTKPGQSA FVLCRLFKKQ DESVEICVEV EPSESSPTSA 180 RSEFALDQTL LDCKPSVSHL AEKVTSNLQE CQTKSSEFTA TEIDQAMQEN FGDVFTKVMS 240 EPLDCTLFPP LHSPFPVEKG SSSTSHSVRN DLTTIDEKGQ LPSESDAYIT EFLDSILNTS 300 DENLFDESDC QWNSFFVSET PSNMTFVKDS GSCSESDAEV AQARQPNPDI TSSLCFKDNI 360 ARASPLKMLR MPEDFRTVTS QNSDNEPSIN FRNSCYLDDF CNADVESCEQ DVFSSVSTTG 420 QFHDAFGGFG DSNNHAVGVA SRNIVATGIR TRMRNPQSLP YREGFVNVKQ GDAPRRLRLQ 480 CKLQVGPVRL QSCMPEEDES KPIVAEGVAD AEKELPVNAA TGAMYELKGD SQLASNENSN 540 ISEEQTTDER TKGRLKSTVL MDKDTSNTFR AGHSNRSFAV LFRAAAVVVM FAILVSTWRC 600 LCLNF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 7e-48 | 8 | 163 | 16 | 169 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 7e-48 | 8 | 163 | 16 | 169 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 7e-48 | 8 | 163 | 16 | 169 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 7e-48 | 8 | 163 | 16 | 169 | NO APICAL MERISTEM PROTEIN |
| 3swm_A | 8e-48 | 8 | 163 | 19 | 172 | NAC domain-containing protein 19 |
| 3swm_B | 8e-48 | 8 | 163 | 19 | 172 | NAC domain-containing protein 19 |
| 3swm_C | 8e-48 | 8 | 163 | 19 | 172 | NAC domain-containing protein 19 |
| 3swm_D | 8e-48 | 8 | 163 | 19 | 172 | NAC domain-containing protein 19 |
| 3swp_A | 8e-48 | 8 | 163 | 19 | 172 | NAC domain-containing protein 19 |
| 3swp_B | 8e-48 | 8 | 163 | 19 | 172 | NAC domain-containing protein 19 |
| 3swp_C | 8e-48 | 8 | 163 | 19 | 172 | NAC domain-containing protein 19 |
| 3swp_D | 8e-48 | 8 | 163 | 19 | 172 | NAC domain-containing protein 19 |
| 4dul_A | 7e-48 | 8 | 163 | 16 | 169 | NAC domain-containing protein 19 |
| 4dul_B | 7e-48 | 8 | 163 | 16 | 169 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Csi.8485 | 0.0 | callus| flower| seed | ||||
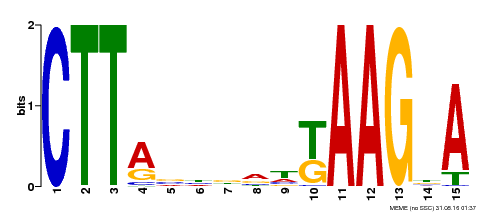
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00398 | DAP | Transfer from AT3G49530 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006478081.1 | 0.0 | NAC domain-containing protein 62-like isoform X2 | ||||
| TrEMBL | A0A067DNB1 | 0.0 | A0A067DNB1_CITSI; Uncharacterized protein | ||||
| STRING | XP_006441279.1 | 0.0 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2416 | 28 | 73 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G49530.1 | 5e-93 | NAC domain containing protein 62 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | orange1.1g007393m |



