 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | orange1.1g005938m | ||||||||
| Common Name | CISIN_1g005938mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 669aa MW: 72332.2 Da PI: 5.5052 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 80.8 | 1.6e-25 | 220 | 270 | 1 | 52 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQk 52
kpr++W+ eLH++Fv+av+qL G++kA+Pk+ilelm+v+gLt+e+v+SHLQ
orange1.1g005938m 220 KPRVVWSVELHQQFVSAVNQL-GIDKAVPKRILELMNVPGLTRENVASHLQE 270
79*******************.***************************974 PP
| |||||||
| 2 | Response_reg | 79.1 | 1.5e-26 | 36 | 144 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHH CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedaleal 90
vl+vdD+ + +++l+q+l++ y +v++++++ al++l+e++ +D++l D+ mp+mdG++ll++i e +lp+i+++a g + + + +
orange1.1g005938m 36 VLVVDDDITCLRILEQMLRRCLY-NVTTCSQAAVALDILRERKgcFDVVLSDVHMPDMDGFKLLEHIGLEM-DLPVIMMSADGRVSAVMRGI 125
89*********************.*******************************************6654.8******************* PP
HTTESEEEESS--HHHHHH CS
Response_reg 91 kaGakdflsKpfdpeelvk 109
+ Ga d+l Kp+ +eel +
orange1.1g005938m 126 RHGACDYLIKPIREEELKN 144
****************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.40.50.2300 | 6.5E-42 | 33 | 168 | No hit | No description |
| SuperFamily | SSF52172 | 9.68E-37 | 33 | 162 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 2.8E-31 | 34 | 146 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 43.303 | 35 | 150 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 1.9E-23 | 36 | 144 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 2.53E-28 | 37 | 149 | No hit | No description |
| SuperFamily | SSF46689 | 1.49E-16 | 217 | 273 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 9.295 | 217 | 276 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-26 | 218 | 282 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.3E-22 | 220 | 271 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 3.8E-6 | 222 | 269 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010380 | Biological Process | regulation of chlorophyll biosynthetic process | ||||
| GO:0031537 | Biological Process | regulation of anthocyanin metabolic process | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0080036 | Biological Process | regulation of cytokinin-activated signaling pathway | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 669 aa Download sequence Send to blast |
MAALQRIVQS SGGSGYGSSR AADVAVPDQF PAGLRVLVVD DDITCLRILE QMLRRCLYNV 60 TTCSQAAVAL DILRERKGCF DVVLSDVHMP DMDGFKLLEH IGLEMDLPVI MMSADGRVSA 120 VMRGIRHGAC DYLIKPIREE ELKNIWQHVV RKRWNENKEH ENSGSLEETD HHKRGSDEIE 180 YASSVNEGTE GTFKAQRKRI SAKEEDDGEL ESDDPSTTKK PRVVWSVELH QQFVSAVNQL 240 GIDKAVPKRI LELMNVPGLT RENVASHLQE INLQKFRLYL KRLNGVSQQG GITNSFCAPI 300 ETNVKLGSLG RFDIQALAAS GQIPPQTLAA LHAELLGRPT GNLVTAVDQP ALLQATLQGP 360 KCIPADHGVA FGQQLVKCQS NVPENLPQSI ISVDDASTGF GVWASNSLGA VASTSNLGGL 420 NPQNGNMLMD ILHQQQQKQQ NQQSQQQSTL SETSRSISVQ PSCLVVPSRS SASFQAGNSP 480 ASVNQSCSFN RGAVVDYSLL SSQSNNSSLN MGQISDGDIK TTAVITGYLA PGSLSPSASS 540 CSVTADNTSS QQVQNSNIAF RAARQVPGLV SNVGDIQGSY GAKSGEVFDL GSLRNHGFMG 600 KGNCIPSRLA VDEFESPMNN LNYGNIYLDN NANKVKEEPN LEFVENAKVG IPMYQQYAPN 660 DLMSVFTD* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 3e-20 | 216 | 284 | 1 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Csi.1503 | 0.0 | flower| stem | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specific DNA sequence. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. May directly activate some type-A response regulators in response to cytokinins. {ECO:0000250|UniProtKB:Q940D0}. | |||||
| UniProt | Transcriptional activator that binds specific DNA sequence. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. May directly activate some type-A response regulators in response to cytokinins. {ECO:0000250|UniProtKB:Q940D0}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
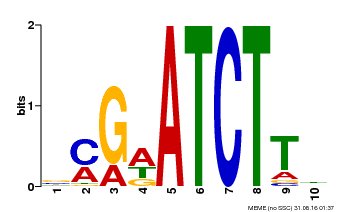
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006469257.1 | 0.0 | two-component response regulator ORR21 isoform X1 | ||||
| Refseq | XP_006469258.1 | 0.0 | two-component response regulator ORR21 isoform X1 | ||||
| Refseq | XP_006469259.1 | 0.0 | two-component response regulator ORR21 isoform X1 | ||||
| Swissprot | A2XE31 | 1e-165 | ORR21_ORYSI; Two-component response regulator ORR21 | ||||
| Swissprot | Q8H7S7 | 1e-165 | ORR21_ORYSJ; Two-component response regulator ORR21 | ||||
| TrEMBL | A0A067FBE5 | 0.0 | A0A067FBE5_CITSI; Two-component response regulator | ||||
| STRING | XP_006469257.1 | 0.0 | (Citrus sinensis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM8150 | 27 | 36 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G16857.1 | 1e-123 | response regulator 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | orange1.1g005938m |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



