 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | orange1.1g004026m | ||||||||
| Common Name | CISIN_1g002869mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 779aa MW: 85071.9 Da PI: 6.0009 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 65.7 | 6.3e-21 | 77 | 132 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l L++rqVk+WFqNrR+++k
orange1.1g004026m 77 KKRYHRHTPQQIQELESLFKECPHPDEKQRLELSKRLCLETRQVKFWFQNRRTQMK 132
688999***********************************************999 PP
| |||||||
| 2 | START | 203.2 | 1.1e-63 | 284 | 509 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE.....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....E CS
START 1 elaeeaaqelvkkalaeepgWvkss.....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....k 78
ela++a++elvk+a+ +ep+W +s + +n +e+l++f++ + + +ea+r++g+v+ ++ lve+l+d + +W e+++ +
orange1.1g004026m 284 ELALAAMDELVKMAQTDEPLWIRSFegsgrQVLNHEEYLRTFTPCIGlkpngFVTEASRETGMVIINSLALVETLMDPN-RWAEMFPcmiaR 374
5899********************99*99999***********99989*9*9***************************.************ PP
EEEEEEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEE CS
START 79 aetlevissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghsk 163
+t++vissg galqlm aelq+lsplvp R++ f+R+++q+ +g+w++vdvS+d ++ + + +v +++lpSg+++++++ng+sk
orange1.1g004026m 375 TATTDVISSGmggtrnGALQLMHAELQVLSPLVPvREVNFLRFCKQHAEGVWAVVDVSIDTIRETSGAPAFVNCRRLPSGCVVQDMPNGYSK 466
****************************************************************999************************* PP
EEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 164 vtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
vtwveh++++++++h+l+++l+ sg+ +ga++wvatlqrqce+
orange1.1g004026m 467 VTWVEHAEYDESQVHQLYKPLIISGMGFGAQRWVATLQRQCEC 509
*****************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.07E-20 | 60 | 134 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.9E-21 | 69 | 134 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.248 | 74 | 134 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.2E-17 | 75 | 138 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.69E-18 | 76 | 134 | No hit | No description |
| Pfam | PF00046 | 1.8E-18 | 77 | 132 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 109 | 132 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 44.451 | 275 | 512 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 3.3E-34 | 277 | 509 | No hit | No description |
| CDD | cd08875 | 1.54E-126 | 279 | 508 | No hit | No description |
| SMART | SM00234 | 2.3E-47 | 284 | 509 | IPR002913 | START domain |
| Pfam | PF01852 | 1.6E-55 | 284 | 509 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 7.14E-24 | 537 | 771 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 779 aa Download sequence Send to blast |
MMIPSVWVCD LVFQQQPNID NQGGGDLQLQ RMGESFEGII GRRSREDLLE HESRSGSDNM 60 DGASGDDLDA ADNPPRKKRY HRHTPQQIQE LESLFKECPH PDEKQRLELS KRLCLETRQV 120 KFWFQNRRTQ MKTQLERHEN SLLRQENDKL RAENMSIRDA MRNPICTNCG GPAIIGDISL 180 EEQHLRIENA RLKDELDRVC ALAGKFLGRP VSSMGPPPMP NSSLELGVGT INGFGGLSST 240 VTTTLPADFG TGISNALPVV MPPNRSGPGV TGLDRSIERS MFLELALAAM DELVKMAQTD 300 EPLWIRSFEG SGRQVLNHEE YLRTFTPCIG LKPNGFVTEA SRETGMVIIN SLALVETLMD 360 PNRWAEMFPC MIARTATTDV ISSGMGGTRN GALQLMHAEL QVLSPLVPVR EVNFLRFCKQ 420 HAEGVWAVVD VSIDTIRETS GAPAFVNCRR LPSGCVVQDM PNGYSKVTWV EHAEYDESQV 480 HQLYKPLIIS GMGFGAQRWV ATLQRQCECL AILMSTSVSA RDHTAITAGG RRSMLKLAQR 540 MTDNFCAGVC ASTVHKWNKL NAGNVDEDVR VMTRKSVDDP GEPPGIVLSA ATSVWLPVSP 600 QRLFNFLRDE RLRSEWDILS NGGPMQEMAH IAKGQDHGNC VSLLRASAIN ANQSSMLILQ 660 ETCTDAAGSL VVYAPVDIPA MHVVMNGGDS AYVALLPSGF AIVPDGPDSR GPLANGPTSG 720 NGSNGGSQRV GGSLLTVAFQ ILVNSLPTAK LTVESVETVN NLISCTVQKI KAALQCES* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Csi.1217 | 0.0 | fruit| vegetative meristem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves and floral buds. {ECO:0000269|PubMed:10402424}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
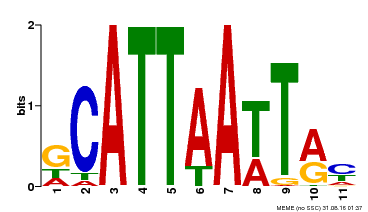
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006445143.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X1 | ||||
| Refseq | XP_006491020.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X1 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A067H2D2 | 0.0 | A0A067H2D2_CITSI; Uncharacterized protein | ||||
| STRING | XP_006491020.1 | 0.0 | (Citrus sinensis) | ||||
| STRING | XP_006445143.1 | 0.0 | (Citrus clementina) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | orange1.1g004026m |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



