 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cucsa.252760.1 | ||||||||
| Common Name | Csa_3G165610, LOC101209013 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 661aa MW: 72810.7 Da PI: 6.0687 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 91.4 | 7.9e-29 | 203 | 256 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv av+qL G++kA+Pk+ilelm+v+gLt+e+v+SHLQkYRl
Cucsa.252760.1 203 KPRVVWSVELHQQFVAAVNQL-GIDKAVPKKILELMNVPGLTRENVASHLQKYRL 256
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 75.9 | 1.5e-25 | 26 | 134 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHHTT CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalealkaG 93
vl+vdD+p+ +++l+++l+ +y +v+ ++ +e+al ll++++ +D++l D+ mp+mdG++ll+ i e +lp+i+++ + ++ + + + G
Cucsa.252760.1 26 VLVVDDDPTCLMILEKMLRICRY-DVTNCSRAEDALSLLRQNKngFDIVLSDVHMPDMDGFKLLEYIGLEM-DLPVIMMSVDDGKNVVMKGVTHG 118
89*********************.***************888889**********************6654.8********************** PP
ESEEEESS--HHHHHH CS
Response_reg 94 akdflsKpfdpeelvk 109
a d+l Kp+ +e+l +
Cucsa.252760.1 119 ACDYLIKPVRMEALKN 134
***********99986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 1.8E-175 | 5 | 651 | IPR017053 | Response regulator B-type, plant |
| Gene3D | G3DSA:3.40.50.2300 | 1.4E-40 | 23 | 162 | No hit | No description |
| SuperFamily | SSF52172 | 8.82E-35 | 23 | 147 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 4.1E-28 | 24 | 136 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 41.122 | 25 | 140 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 1.2E-22 | 26 | 134 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 4.96E-25 | 27 | 139 | No hit | No description |
| SuperFamily | SSF46689 | 3.58E-20 | 200 | 260 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 11.048 | 200 | 259 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-30 | 202 | 261 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 9.4E-25 | 203 | 256 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 9.4E-8 | 205 | 255 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0016310 | Biological Process | phosphorylation | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0016301 | Molecular Function | kinase activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 661 aa Download sequence Send to blast |
MSTASSSSVR KAGEAVPDQF PAGLRVLVVD DDPTCLMILE KMLRICRYDV TNCSRAEDAL 60 SLLRQNKNGF DIVLSDVHMP DMDGFKLLEY IGLEMDLPVI MMSVDDGKNV VMKGVTHGAC 120 DYLIKPVRME ALKNIWQHVV RKRKNEWKDL EQTCVDDVDR QQKTNEDADY SSSANEGSWR 180 NSKRRKDDVE DPEERDDSST LKKPRVVWSV ELHQQFVAAV NQLGIDKAVP KKILELMNVP 240 GLTRENVASH LQKYRLYLRR LSGITQHQSN LNNTFMSAQD AFGPPLNGLD LQTLAAAGQL 300 QPQSLATLQA AGFGRSTAKS GMPMPLVDQR NHIFSFENPK LRFGDGQQPH LNGSKPMNLL 360 HGIPTTMEPK QLANLQHSAQ SHGNMTMQVS IQGGQSSSQL MQTPQPQARA QILNESTTTS 420 VTRLPQTMQP SILPNGTTSA VLARTEFGNN NRGGGYNLVS PASTMLNFPL NQTAELPGNS 480 FPLQSTPGMS SIVPKGRFPD DVNSDIKGSE GFGPSYDMFR DLHPQKPHDW DLQHVGVTFD 540 TSQGSLDIPP SAFSHQGYAS SQQNGQNRNT STAGKAMFLL EEGSDNGNAQ SMGQQLNPIF 600 VDGSVRVKSE RASDISSQTD LFSEPFGQED LMSSLFKQQQ GSIATAESEL EWLFHHNTPV 660 * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 3e-22 | 202 | 262 | 4 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]GATT-3'. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. Could directly activate some type-A response regulators in response to cytokinins. Involved in the expression of nuclear genes for components of mitochondrial complex I. Promotes cytokinin-mediated leaf longevity. Involved in the ethylene signaling pathway in an ETR1-dependent manner and in the cytokinin signaling pathway. {ECO:0000269|PubMed:11370868, ECO:0000269|PubMed:11574878, ECO:0000269|PubMed:15282545, ECO:0000269|PubMed:16407152}. | |||||
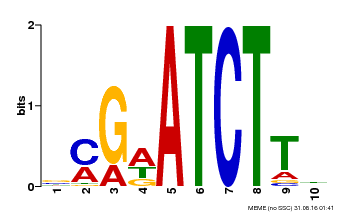
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681848 | 0.0 | LN681848.1 Cucumis melo genomic scaffold, anchoredscaffold00006. | |||
| GenBank | LN713260 | 0.0 | LN713260.1 Cucumis melo genomic chromosome, chr_6. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004134232.1 | 0.0 | PREDICTED: two-component response regulator ARR2 | ||||
| Swissprot | Q9ZWJ9 | 0.0 | ARR2_ARATH; Two-component response regulator ARR2 | ||||
| TrEMBL | A0A0A0L800 | 0.0 | A0A0A0L800_CUCSA; Two-component response regulator | ||||
| STRING | XP_004134232.1 | 0.0 | (Cucumis sativus) | ||||
| STRING | XP_004162202.1 | 0.0 | (Cucumis sativus) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3958 | 34 | 58 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 0.0 | response regulator 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cucsa.252760.1 |
| Entrez Gene | 101209013 |



