 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cucsa.180090.2 | ||||||||
| Common Name | Csa_4G056770, LOC101210541 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 809aa MW: 94208 Da PI: 8.0826 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 87.1 | 2.6e-27 | 47 | 133 | 2 | 91 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHelap 91
fY+eYAk++GF++++++s++sk+++e+++++f Cs++g + e+++ ++r+ ++++t+Cka+++vk++ dg+w ++++ ++HnHel p
Cucsa.180090.2 47 FYQEYAKSMGFTTSIKNSRRSKKSKEFIDAKFACSRYGVTPESESG---NSRRPSVKKTDCKASMHVKRRPDGRWIIHEFIKDHNHELLP 133
***************************************9998877...7788899*******************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 5.8E-25 | 47 | 133 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 8.3E-25 | 253 | 346 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.371 | 533 | 569 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 1.3E-5 | 542 | 568 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.3E-7 | 544 | 571 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 809 aa Download sequence Send to blast |
MVDVVAEMQD RGGIVSLPKK DILFEGDVDF EPHTGIEFES HEAAYTFYQE YAKSMGFTTS 60 IKNSRRSKKS KEFIDAKFAC SRYGVTPESE SGNSRRPSVK KTDCKASMHV KRRPDGRWII 120 HEFIKDHNHE LLPALAYHFR IHRNVKLAEK NNIDILHAVS ERTRRMYVEM SKQCGGYRNF 180 SFPQIDTTYQ FDKGRYLALD EGDAQMLLEY FKRVQKENPY FFYAIDLNEE QRLRNLFWVD 240 AKSRNDYVSF SDVVSFDISY IKTNDKLPFA PFIGANHHAQ SMVLGCALAA DWTKPTFAWL 300 LKTWLRAMGG KAPKVIITDQ DKALKLAIEE VFPNTRHCFA LWHILEKIPE TLAHVIKRHE 360 NFLAKFNKCI FKSWSDEQFD MRWWKMVTRF ELQDDEWIQS LYGDRKKWVP TYMEDIFLAG 420 MSTTQRSDSM NAFFDKYIHK KITLKEFLRQ YGIILQNRYE EEVIADFDTL HKQPALKSPS 480 PWEKQMSTLY THTIFKKFQV EVLGVVGCRM RKEIEDGTIT TFRVQDCEKD EHFLVRWHKL 540 NSEVSCFCRL FEYKGFLCRH ALIVLQMLDF RSIPSQYILK RWTKDAKSRQ PVTEETEFRQ 600 NRVQRYNDLC KKAIELSEEG SHSEECYNIA IRTLVEALKN CVNINNSKSA PADSCVHAHG 660 LREEEENQGS ITAKANKKKS TNRKRKVQTE TDMILVEAQD NLQPMDSLTS DSMNLTGYYG 720 TQQNVQGLVQ LNLMEPPHDA SYYVSQQSIQ GLGQLNTIAA NHDGFFGVQH NSIHTLVDYR 780 PTTSYSYSLQ EEQHLRSAQL HGSTSRHT* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
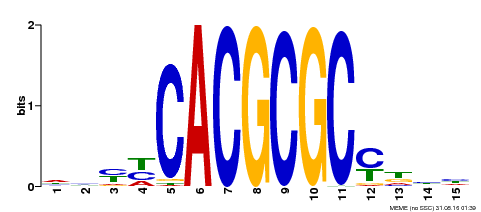
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681873 | 0.0 | LN681873.1 Cucumis melo genomic scaffold, anchoredscaffold00027. | |||
| GenBank | LN713261 | 0.0 | LN713261.1 Cucumis melo genomic chromosome, chr_7. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004147732.1 | 0.0 | PREDICTED: protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| Refseq | XP_011653253.1 | 0.0 | PREDICTED: protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| Refseq | XP_011653254.1 | 0.0 | PREDICTED: protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| Refseq | XP_011653255.1 | 0.0 | PREDICTED: protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A0A0KXC8 | 0.0 | A0A0A0KXC8_CUCSA; Uncharacterized protein | ||||
| STRING | XP_004147732.1 | 0.0 | (Cucumis sativus) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cucsa.180090.2 |
| Entrez Gene | 101210541 |



