 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cucsa.180080.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 816aa MW: 93750.5 Da PI: 7.1562 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 85.5 | 8.5e-27 | 88 | 190 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk............tekerrtraetrtgCkaklkvkkekdgkwevtklel 83
+fY+eYA+++GF++ +++s++sk+++e+++++f Cs++g ++e +k+ +e++ +ra +t+Cka+++vk++ dgkw+++++++
Cucsa.180080.2 88 SFYQEYARSMGFNTAIQNSRRSKTSREFIDAKFACSRYGMKREYDKSfnrprvrqtkqeSENSTGRRACAKTDCKASMHVKRRADGKWVIHSFVK 182
5*******************************************999*****99976555555559999************************** PP
FAR1 84 eHnHelap 91
eHnHel p
Cucsa.180080.2 183 EHNHELLP 190
*****975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 9.1E-25 | 88 | 190 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 5.9E-27 | 288 | 380 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 8.963 | 568 | 604 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 1.3E-5 | 569 | 603 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.6E-7 | 579 | 606 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 816 aa Download sequence Send to blast |
MDIDLRLPSG EHDKEEEPNG INNMLDVEEK LHNGVIESGD MVDATNGMHV EDGGNLNSPM 60 LDMVMFKEDT NLEPLPGMEF ESHSEAYSFY QEYARSMGFN TAIQNSRRSK TSREFIDAKF 120 ACSRYGMKRE YDKSFNRPRV RQTKQESENS TGRRACAKTD CKASMHVKRR ADGKWVIHSF 180 VKEHNHELLP AQAVSEQTRK MYAAMARQFA EYKNVVGLKN DPKNPFDKVR NLAFDAADAK 240 ILLDFLTQMQ NLNSNFFYAV DIGDDHRLRN LFWIDAKSRH DYSYFNDVVS LDTTYIRNKY 300 KLPLAFFVGV NQHYQFMLLG CALLSDETPT TYAWLLHIWL KAIGGQAPKV IITDHDKVLK 360 TAVQEVLPNA YHHFTLWHIL GKFSENLGNI IKRHENFMAK FEKCIYKSWT IEEFEKRWLK 420 LVDRFELKED ELVQSLCEDQ RHWAPTYMKD VFLAGMSMPQ RSESVNSFLD KYLHKKTSVQ 480 EFVKQYETIL QDRYEEEAKA DSDTWNKQPT LRSPSPFEKS VSGLYTHAVF KKFQVEVLGA 540 VACFPRKVKE DEKNITYKVQ DLEKDLEFVV VWNGLKSEVS CLCRLYEYKG YLCRHAMVVL 600 QKCELSTIPA QYILKRWTKD AKSRQLMGEE LEPVQSRVQR YNDLCQRALR LIEEGSMSQE 660 SYSIAVHALE ETLGNCISVN NSNRTFLEAG TSAAHGLLCI EEDSHIRSIG KTNKKKNPTK 720 KRKVNCEPDV MTVGAQDSLQ QMDKLSSRAV TLDGYFGAQP SVQGMVQLNL MAPTRDNYYG 780 NQQAIQGLGQ LNSIAPSHDG YYAAQQSIHG LTCLI* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
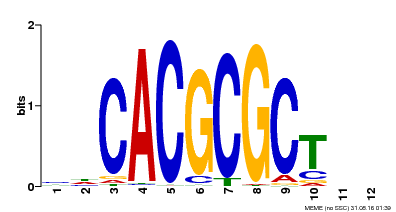
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681873 | 0.0 | LN681873.1 Cucumis melo genomic scaffold, anchoredscaffold00027. | |||
| GenBank | LN713261 | 0.0 | LN713261.1 Cucumis melo genomic chromosome, chr_7. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004147733.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Refseq | XP_011653256.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Refseq | XP_011653257.1 | 0.0 | PREDICTED: protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A1S4DYW2 | 0.0 | A0A1S4DYW2_CUCME; protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| STRING | XP_004147733.1 | 0.0 | (Cucumis sativus) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cucsa.180080.2 |



