 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cucsa.159750.1 | ||||||||
| Common Name | Csa_2G365700, LOC101207697 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | BBR-BPC | ||||||||
| Protein Properties | Length: 339aa MW: 37771.9 Da PI: 10.023 | ||||||||
| Description | BBR-BPC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GAGA_bind | 392.4 | 6.7e-120 | 1 | 338 | 1 | 301 |
GAGA_bind 1 mdddgsre..rnkg.yyepa..aslkenl..glqlmssiaerdakirernlalsekkaavaerdmaflqrdkalaernkalverdnkllalllve 88
mdd+g+re r+k+ +y++a + + +++ +q+m+++aerda+i+ernlalsekkaa+aerdma+lqrd+a+aern+al+erdn++++l+++e
Cucsa.159750.1 1 MDDSGHREngRHKPdQYKSAqgQWMMQHQpsMKQIMAIMAERDAAIQERNLALSEKKAALAERDMAYLQRDAAIAERNNALLERDNAIATLQYRE 95
9*****999*********6656445545422469************************************************************* PP
GAGA_bind 89 nsla..salpvgvqvlsgtksidslqq.lse..pqledsavelreeekleal....pieeaaeeakekkkkkkrqrakkpkekkakkkkkkseks 174
ns++ ++p+g+q+ +g+k+i++ qq +++ p+++++ ++ re+ ++ + p+ ++ ++a+++k++k++++ +p +k +k +
Cucsa.159750.1 96 NSINnnLSCPPGCQIARGVKHIHHPQQqHTHhvPHMNENNYNSREMLASNDPcptsPVASESTKARRNKRPKEGKTVPTPNKKVSKGP------- 183
*9987789****************9994334799********8887654432111144444444455555555555555554444433....... PP
GAGA_bind 175 kkkvkkesader.............................skaekksidlvlngvslDestlPvPvCsCtGalrqCYkWGnGGWqSaCCtttiS 240
+kvk+e +d + sk+++k +dl+ln+v++Dest+P+P+CsCtG+ rqCYkWGnGGWqSaCCttt+S
Cucsa.159750.1 184 -RKVKREAEDLNkimlgksqewkdgigimsagddlnkqlvvSKSDWKGQDLGLNQVAFDESTMPAPICSCTGVIRQCYKWGNGGWQSACCTTTLS 277
.33333333322244444556889*********************************************************************** PP
GAGA_bind 241 vyPLPvstkrrgaRiagrKmSqgafkklLekLaaeGydlsnpvDLkdhWAkHGtnkfvtir 301
+yPLP+++++r+aR++grKmS++af+klL++LaaeG+dls pvDLk+hWAkHGtn+++ti+
Cucsa.159750.1 278 MYPLPAVPNKRHARLGGRKMSGSAFNKLLSRLAAEGHDLSAPVDLKNHWAKHGTNRYITIK 338
************************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF06217 | 3.2E-104 | 1 | 338 | IPR010409 | GAGA-binding transcriptional activator |
| SMART | SM01226 | 2.2E-175 | 1 | 338 | IPR010409 | GAGA-binding transcriptional activator |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 339 aa Download sequence Send to blast |
MDDSGHRENG RHKPDQYKSA QGQWMMQHQP SMKQIMAIMA ERDAAIQERN LALSEKKAAL 60 AERDMAYLQR DAAIAERNNA LLERDNAIAT LQYRENSINN NLSCPPGCQI ARGVKHIHHP 120 QQQHTHHVPH MNENNYNSRE MLASNDPCPT SPVASESTKA RRNKRPKEGK TVPTPNKKVS 180 KGPRKVKREA EDLNKIMLGK SQEWKDGIGI MSAGDDLNKQ LVVSKSDWKG QDLGLNQVAF 240 DESTMPAPIC SCTGVIRQCY KWGNGGWQSA CCTTTLSMYP LPAVPNKRHA RLGGRKMSGS 300 AFNKLLSRLA AEGHDLSAPV DLKNHWAKHG TNRYITIK* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds to GA-rich elements (GAGA-repeats) present in regulatory sequences of genes involved in developmental processes. {ECO:0000269|PubMed:14731261}. | |||||
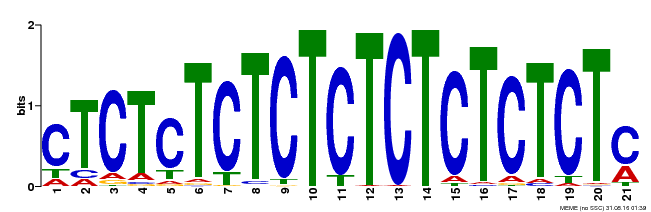
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00540 | DAP | Transfer from AT5G42520 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681823 | 0.0 | LN681823.1 Cucumis melo genomic scaffold, anchoredscaffold00014. | |||
| GenBank | LN713257 | 0.0 | LN713257.1 Cucumis melo genomic chromosome, chr_3. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004152657.1 | 0.0 | PREDICTED: protein BASIC PENTACYSTEINE6 | ||||
| Swissprot | Q8L999 | 1e-152 | BPC6_ARATH; Protein BASIC PENTACYSTEINE6 | ||||
| TrEMBL | A0A0A0LLL3 | 0.0 | A0A0A0LLL3_CUCSA; Uncharacterized protein | ||||
| STRING | XP_004165561.1 | 0.0 | (Cucumis sativus) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF6905 | 34 | 49 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G42520.1 | 1e-144 | basic pentacysteine 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cucsa.159750.1 |
| Entrez Gene | 101207697 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



