 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | estExt_fgenesh1_pg.C_50154 | ||||||||
| Common Name | COCSUDRAFT_46871 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Trebouxiophyceae; Trebouxiophyceae incertae sedis; Coccomyxaceae; Coccomyxa; Coccomyxa subellipsoidea
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 405aa MW: 42780.3 Da PI: 9.8478 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 51.6 | 1.3e-16 | 62 | 90 | 1 | 29 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGly 29
Cs+Cgt Tp+WR+gp+g+ktLCnaCG++
estExt_fgenesh1_pg.C_50154 62 CSQCGTNRTPQWREGPEGPKTLCNACGVK 90
***************************87 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50114 | 12.057 | 56 | 89 | IPR000679 | Zinc finger, GATA-type |
| SMART | SM00401 | 2.0E-8 | 56 | 106 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 3.8E-11 | 59 | 93 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 9.9E-13 | 60 | 89 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 4.39E-12 | 61 | 96 | No hit | No description |
| PROSITE pattern | PS00344 | 0 | 62 | 87 | IPR000679 | Zinc finger, GATA-type |
| Pfam | PF00320 | 6.0E-14 | 62 | 90 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 405 aa Download sequence Send to blast |
MAVVQQYMAA AEMGVAAAYL MEMGGNGGKI TSKEVFKQAA ARSSLPYARP NFIGLGAGGK 60 TCSQCGTNRT PQWREGPEGP KTLCNACGVK RVRQMRMLTD GHKRRPPAAA TAPLKTALSL 120 AKQSSGSLTS DSNAVIRRRD RSPAVRRPQR KAAAKAASRT AEFATTGDWP DDEADASYPV 180 RSAAQSTFTD PSGSEDSAAL NSDSAEEVAW TPTRGPGPAH TSMPQPPSPF PVVGHETAAA 240 VNLLSMSIHE QDLGAGRAAA GALAARDLST FYEHHQYGKQ MGSPMTFNSA AFAAMVNSSM 300 NEVPGAKPEE QPAAAEELKD ALRRLPHGKQ TEMALVKERL ENSLREEAAA NAAVAAVAKI 360 LAMKQAAGIR ARAAAAAASR DMQRFMAELD EEFSLSNILK KQRT* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00554 | DAP | Transfer from AT5G49300 | Download |
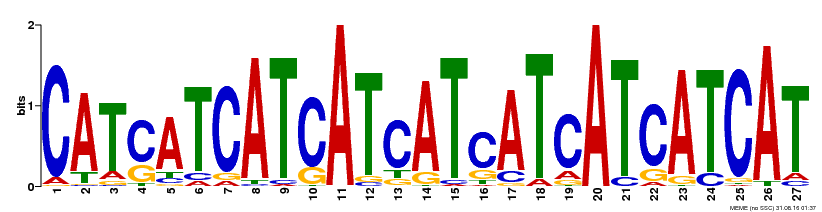
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_005649143.1 | 0.0 | hypothetical protein COCSUDRAFT_46871 | ||||
| TrEMBL | I0Z1T0 | 0.0 | I0Z1T0_COCSC; Uncharacterized protein | ||||
| STRING | XP_005649143.1 | 0.0 | (Coccomyxa subellipsoidea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP220 | 14 | 30 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G32890.1 | 2e-12 | GATA transcription factor 9 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 17042601 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



