 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | estExt_Genemark1.C_180179 | ||||||||
| Common Name | COCSUDRAFT_67762 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Trebouxiophyceae; Trebouxiophyceae incertae sedis; Coccomyxaceae; Coccomyxa; Coccomyxa subellipsoidea
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 1035aa MW: 109686 Da PI: 5.1627 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 114 | 8.6e-36 | 77 | 153 | 1 | 77 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S-- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkq 77
+Cqv+gC+a+l+ +keyh+r+k+Ce+h k ++++ +g++qrfCqqC+rfh+l+ fD +krsCr rL++hn+rrrkk
estExt_Genemark1.C_180179 77 VCQVDGCSAELTGLKEYHQRYKICEFHLKVNSIIREGKRQRFCQQCGRFHDLTSFDGDKRSCRARLQRHNARRRKKP 153
6*************************************************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 6.5E-30 | 72 | 139 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 28.462 | 75 | 152 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 3.01E-32 | 76 | 155 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 8.6E-31 | 78 | 152 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 2.1E-6 | 668 | 819 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.87E-7 | 707 | 823 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 9.49 | 708 | 817 | IPR020683 | Ankyrin repeat-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010321 | Biological Process | regulation of vegetative phase change | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1035 aa Download sequence Send to blast |
MAENALDAKE RLWNPQDWCW DPYNMLALAS DRKGLAVAGK AVFGGLPYAV QPQLGPAASL 60 LLDDVNRSGC RGKGPAVCQV DGCSAELTGL KEYHQRYKIC EFHLKVNSII REGKRQRFCQ 120 QCGRFHDLTS FDGDKRSCRA RLQRHNARRR KKPAESGGSG AKPGVKKPPP TPLMVKGSRD 180 FVRTDSSKQR QTSDSTMDQM STPQGSLEAS SRQQMSIKSG SMESQGTIRQ LIRGGRTLPE 240 EGPSGQQLAA RLQSSSFASE QMHVTGFGDS DGAPMTAGQS SVPSSNYGDI PPSTYAADSS 300 MENLDCLQDI NFFAPQQSSP MPDYSDLNQA ATQMQATFAA DPAAMDLGQA AAASKDTTAF 360 LVEEARALLS YAGQYQALEY TAEEHFTRMS AKMFKCTPEQ LPADLKANLL AMLQCGVNAL 420 EGYIRPGCVH LTLHAMLGPD AGAALQAGCH SGMGVRALVH NLVTASNDTL WRSETLLIQW 480 YEEVALVRYG KVLHVLSASG SRGVLPRLHS VSPLAVTSAA ATTVKLIGSN IAAPDNTVLA 540 QSQGENVAIE VVSAEADMEG APEEVSSMLE VAVPAGQRNG VVHLEVARGA LLSGPKPLLI 600 LKDAAAAEEI NQLAQGDSTG LDVDGFLLDM GRVLELKNAV AAAKFGTEVH VRTFAADDVA 660 VIAQIARRQL IVAINKGWLS VARLVLPVAM ADAKSVAEAV RKLEELTDGL PLLHLAVRSQ 720 SASLVELLLE WTSVGGYELR AGTPGPHGLT ALHLAAMLKD GGKLAALLTA LCVDGATAWT 780 QAGSAGASPA DLAIAMGNHG THELVYQQLC DTMAASTKGS SDEELEDADM CDDEDYDDIE 840 VDSETGELLD LSEIICRVRG GGSDVDTLDA ALHQHTKNGE PAHVEAAAKA QPEAARTATG 900 TAEPPQHAPI HKVSPDEAGR SDGSSYTSTS RSSSDANSLL SDWPSQSCDS LGLRKRSPDV 960 PPVREGVEFT PEQKGGSKGR DADYSSLYNI GSCIGLKVAP TVFDRTSAIG SALLIAACGA 1020 LTLFFRYFGS IFLD* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 4e-25 | 74 | 151 | 7 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00060 | PBM | Transfer from AT3G60030 | Download |
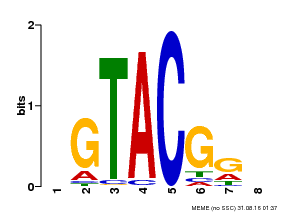
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_005644255.1 | 0.0 | hypothetical protein COCSUDRAFT_67762 | ||||
| TrEMBL | I0YMU2 | 0.0 | I0YMU2_COCSC; Uncharacterized protein | ||||
| STRING | XP_005644255.1 | 0.0 | (Coccomyxa subellipsoidea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP5500 | 5 | 8 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G60030.1 | 8e-25 | squamosa promoter-binding protein-like 12 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 17037683 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



