 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | estExt_Genemark1.C_110100 | ||||||||
| Common Name | COCSUDRAFT_66702 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Trebouxiophyceae; Trebouxiophyceae incertae sedis; Coccomyxaceae; Coccomyxa; Coccomyxa subellipsoidea
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 728aa MW: 77208.2 Da PI: 6.1064 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 73.9 | 2e-23 | 357 | 406 | 3 | 52 |
RWP-RK 3 keisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
k+++ledl+++F +k+AA+ Lg+c+T+LKr+CR++GI+RWP+R++ +l
estExt_Genemark1.C_110100 357 KRLRLEDLQSQFGVGLKEAANRLGICPTTLKRACRRHGIQRWPRRQLLKL 406
7899******************************************9875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 16.222 | 334 | 428 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 3.4E-20 | 359 | 405 | IPR003035 | RWP-RK domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 728 aa Download sequence Send to blast |
MVNTEVYDLP HSVEQRGAMA SRIARAVLNF QDQLLEADSN TGSLCQVWLP EVSEDGDVVL 60 RTKGLPFCVA GVGDLLALFR CISCRYCFGT DATKPDMLGA PGRVFTTQEP EMSRNVQKYS 120 KEVYLRASEA QQCRVHSTVV LPLFTSADRK SSLGVVEVVQ TRQDMSFAEI VSNLARSLED 180 CNLFTCEGLR GKEMAESSDA IKQMVKGGLV PSSSTRSNSS EADKAAATGD AENEGTSGLT 240 GRASTASISV DLADAASLEA ERKKSLGTRT RNASLAADLR AALRQQSLRD SAPKEALAAA 300 AAAPTAGAGA EQMSARGGSL SEEEVDMDED DDLEDETEEE RARRKGKGAG NPGKPGKRLR 360 LEDLQSQFGV GLKEAANRLG ICPTTLKRAC RRHGIQRWPR RQLLKLSRAI DQINATGSVK 420 SADGSSPAAN GLQPLPGPDT RWTALAQLIP GIAVQPDNRQ PHKTLPSTQA LALTHSTDQR 480 MLQRVDSSAS VDPYVAAAAA EQLQRYHSNG SLQQGSPVGA ASAGYGLPSH ASAQHLHRHD 540 SGAVAGADAL AGDGSVHRGQ AAFQYLAAAG PSVDAHMQQA QHYGSANSTP MPIPAGSAAH 600 LAHSYPGAMP VGTQPAMMPQ VFDGGQQISA PLSMPAMSLS HVHPPTGQHQ SPPMMVGGEG 660 SYRGQDLLSY SNQRTMIDAK MDEHTHNWMQ HAGNGAQHHL DLFPSVDDLD DDVGLVDAEV 720 LEMLLSR* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
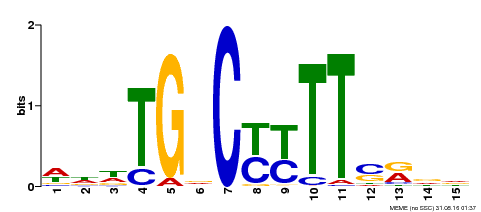
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_005646373.1 | 0.0 | hypothetical protein COCSUDRAFT_66702 | ||||
| TrEMBL | I0YTW0 | 0.0 | I0YTW0_COCSC; Uncharacterized protein | ||||
| STRING | XP_005646373.1 | 0.0 | (Coccomyxa subellipsoidea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP3760 | 5 | 6 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 3e-20 | NIN like protein 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 17039814 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



