 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Genemark1.2565_g | ||||||||
| Common Name | COCSUDRAFT_61775 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Trebouxiophyceae; Trebouxiophyceae incertae sedis; Coccomyxaceae; Coccomyxa; Coccomyxa subellipsoidea
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1550aa MW: 165677 Da PI: 8.1256 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 145.4 | 1.5e-45 | 95 | 216 | 2 | 116 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltl..elktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahs.... 88
+k ++ wlkn+e++++L ++++++l ++++ p+ gsl+L++r+ vr+frkDG++w+kk dgktvrE+hekLKvg+ve+l+cyYah+
Genemark1.2565_g 95 HKAQSAWLKNTEVCDLLLHYAEYNLPVarDPPNLPPGGSLFLFDRRAVRFFRKDGHNWRKKADGKTVRETHEKLKVGNVEMLNCYYAHAdtee 187
55699******************9876679**********************************************************96666 PP
CG-1 89 ..eenptfqrrcywlLeeelekivlvhyle 116
++ + +qrrcywlLe+e ++ivlvhyl+
Genemark1.2565_g 188 gaQQATRLQRRCYWLLESE-DDIVLVHYLN 216
6567789************.********98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 53.367 | 90 | 223 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.8E-53 | 93 | 218 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 3.4E-44 | 97 | 216 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:1.25.40.20 | 1.3E-12 | 692 | 719 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF81296 | 2.96E-14 | 837 | 922 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 3.59E-11 | 1051 | 1152 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.3E-12 | 1053 | 1152 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 8.03E-10 | 1054 | 1163 | No hit | No description |
| PROSITE profile | PS50297 | 12.063 | 1059 | 1162 | IPR020683 | Ankyrin repeat-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1550 aa Download sequence Send to blast |
MQPLEGPLAR MDSDADVKLI IAMDKGGCEA NVVLRRDAQQ EDILKAYVQG YMLLHSNAPM 60 GQQPQRLPPG TAKLPAQSLS PDNPRFPQKV RDILHKAQSA WLKNTEVCDL LLHYAEYNLP 120 VARDPPNLPP GGSLFLFDRR AVRFFRKDGH NWRKKADGKT VRETHEKLKV GNVEMLNCYY 180 AHADTEEGAQ QATRLQRRCY WLLESEDDIV LVHYLNIDKA KGKTEDSWAP SDPEMGRRPG 240 FTVTSGGVHG MPGLGIQGLA GHPAHQLLQQ HHHQQHPVDP YIPLMPSLSL DSFFGAYDDR 300 DAAQRAQAPP PMMENVLPAW EDLSNGPSNS LQAAADWRSM LRGDSLSMGL SEGPTAALVE 360 HGANMMMIEQ GAQQPETWAQ DIPVAMRTPS LEARQARAAA VMRQNAELQQ RQSSLERRAA 420 YQKSLLAANR DQMQARPQSP LLFRQHQAAA AAMLQQAQQG GQANGMPYNL NAFQQRLLAA 480 ARQGPPGAGG GPMHGQMPGI GFRPQEPQAA AQAAALAGLA GQMQPPHYNQ QMPPPPPPQR 540 PFMAANTSGA SAAPGVGMPQ ARLPPLQQGA YQGGPAAGPV PADWARAYRE QQHKAAQAAI 600 ARSAKWGTTD PPLRPVPLRQ GSAPGPLDIS PSGSAGPSPG KPPLAATEGF PKMSKLDALL 660 MQQQQQQQQH SGSSSGEGKQ TMSSPGALGA HASASGDQIL HHASSAGQLG AIGTQVKMDE 720 TASEGTVTGT PLGRAERPAS KVAKALALEA QLRSVDGAVM EQLQTARAES GCSASLDSKV 780 QKLEEDLDHL EASSRQLLQD SVQGPGMEAA GVAMQEPATS GTHTSSLSHA PSASLELLDF 840 SPEWDFTLGG TKVIVTCREV DGDITSNCPV CVMFDKEQVP AARLQAGVYR CHAPPHEAGT 900 VGLCVTYGDG RPRSNVQPFT YRGTPLTARA QDDLARAAIP DRDLQLRLIH MLMSSSKGAT 960 SSSTVSPASP SNSSDSNTHK QHASPSRTAA PTAGSATVEV ALEDNPNALQ YLSDDLREKL 1020 LQTLLERRLK QFTSDVREGK AQQGSGWSPS FAVNRRAQSG LALVHILAAL GYDWGLQLLI 1080 PLGALLDLQD AWGRTALHWA ATYACEATVV LLLVRCAHPA PLSHGGEAQP RATPADMAAG 1140 NGHAGIAAFL SEQALLRLAK DNNVSLDEPA DSRTGDQSMS ALRKRARSLR RADSASELPL 1200 LREAMAAPDG ASKFPEEEGD EGTSPKEVLA RVKKLAERVR QALTRAACRR AAMEQQSLRR 1260 HKKEGDAVAR IESSLQAWQA VQQAVQRGIA QSDSDAAARR LVRLGERISC INACLRDAIK 1320 RSLGCGGASS ALAPVDEAAA LPITGNSMVL DYGSDSDAHA MAGDSLLRRR SLIRLRESRR 1380 RDSITKIKGF ESDTSIRQFL ARGREGSKAD EGGPSQLDIR DLRSLSLSFP SFSVVGPPIG 1440 VSLPDCDLRE LRNFIAGASA KVHRSAFGTL ASASNAKAAA APARAGAAPE ASAEMALVQK 1500 AVAHIEALQE EGPGHAQYLR LCQAYTQLTT TQPWGTFKQA RRGSLSNAV* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1181 | 1187 | LRKRARS |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
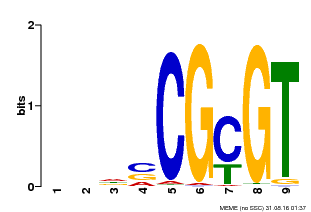
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_005650122.1 | 0.0 | hypothetical protein COCSUDRAFT_61775 | ||||
| TrEMBL | I0Z4K9 | 0.0 | I0Z4K9_COCSC; Uncharacterized protein | ||||
| STRING | XP_005650122.1 | 0.0 | (Coccomyxa subellipsoidea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP5132 | 8 | 9 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 7e-41 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 17043580 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



