 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PK16974.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Cannabis
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 823aa MW: 91686 Da PI: 6.8961 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 74.9 | 9.2e-24 | 150 | 251 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvv 94
f+k+lt sd++++g +++ +++a+e+ + + + +++l+ +d++g++W++++i+r++++r++l++GW+ Fv++++L +gD ++F + +++el+v
PK16974.1 150 FCKTLTASDTSTHGGFSVLRRHADEClppldmSRQ-PPTQELVAKDLHGNEWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL--RGENGELRV 246
99*****************************7333.34459************************************************..449999*** PP
EEE-S CS
B3 95 kvfrk 99
+v+r+
PK16974.1 247 GVRRA 251
**996 PP
| |||||||
| 2 | Auxin_resp | 120 | 1.8e-39 | 276 | 358 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha+st+++F+v+Y+Pr+s++eF+v+++++++a+k+++s+GmRfkm+fe+e+++e+r++Gt++g++++dp rW++SkWr+Lk
PK16974.1 276 AWHAISTGTMFTVYYKPRTSPAEFIVPCDQYLEATKNNYSIGMRFKMKFEGEEAPEQRFTGTIIGIEEFDPNRWKDSKWRCLK 358
89*******************************************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 1.4E-41 | 146 | 265 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 4.71E-40 | 146 | 279 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 7.26E-22 | 148 | 250 | No hit | No description |
| Pfam | PF02362 | 1.1E-21 | 150 | 251 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 2.0E-23 | 150 | 252 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 12.012 | 150 | 252 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 2.0E-36 | 276 | 358 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 3.9E-6 | 681 | 734 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 3.73E-11 | 694 | 775 | No hit | No description |
| PROSITE profile | PS51745 | 23.809 | 698 | 782 | IPR000270 | PB1 domain |
| Pfam | PF02309 | 2.1E-6 | 744 | 787 | IPR033389 | AUX/IAA domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 823 aa Download sequence Send to blast |
MALSEASVKG NSGNPMGESF SSSFTDHNDA RKAAEARKDA EAALYEELWH ACAGPLVTVP 60 RKGERVFYFP QGHIEQVEAS TNQVADQQMP VYNLPPRILC RVMNVELKAE ADTDEVFAQI 120 TLMPDPNQND NAFEKETPPP PPPRFQVHSF CKTLTASDTS THGGFSVLRR HADECLPPLD 180 MSRQPPTQEL VAKDLHGNEW RFRHIFRGQP RRHLLQSGWS VFVSSKRLVA GDAFIFLRGE 240 NGELRVGVRR AMRKQENVPS SVISSHSMHL GVLATAWHAI STGTMFTVYY KPRTSPAEFI 300 VPCDQYLEAT KNNYSIGMRF KMKFEGEEAP EQRFTGTIIG IEEFDPNRWK DSKWRCLKVR 360 WDETSAIPRP ERVSPWKIEP ALAPPALNPL PVPRSKRPRS NIVPLSPDSS VLTREGSSKV 420 TVDPSLPSGF SRVLQGQELS TLRGNFVESN DSDTAEKSVV WQPSLDDEKV DVVSASRRYV 480 PENRASSGKH EPTYTDLLSG FGTTVDSTRG FCSFTEHSVA TANSMRKHSL DQEGRLNLHS 540 SPRSMVPSSL SLSLDTNMKG SVQGAISYQG QGNVRYGGFD DYTTLSGHRV EHAHGNWFMP 600 PPSSPHFENT VHSRELTSKP ISEQKFEVAK PKDGNCKIFG YSLIAPEPAV SHKSSVAKTM 660 GTKIADNPIV SSDPEKPMQT TQQQSREHQG KTQNSSTRSC TKVHKQGIAL GRSVDLTKFN 720 NYKELIAELD ELFEFGGELM APKKNWLIVY TDDENDMMLV GDDPWQEFCG MVKKIFIYTR 780 EEVQKMSPGT LNGHGDGNPS VAEVMDAKEK SQTLPLSSVP ENC |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-178 | 43 | 380 | 18 | 354 | Auxin response factor 1 |
| 4ldw_A | 1e-178 | 43 | 380 | 18 | 354 | Auxin response factor 1 |
| 4ldw_B | 1e-178 | 43 | 380 | 18 | 354 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Could act as transcriptional activator or repressor. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Promotes flowering, stamen development, floral organ abscission and fruit dehiscence. Functions independently of ethylene and cytokinin response pathways. May act as a repressor of cell division and organ growth. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:15960614, ECO:0000269|PubMed:16176952, ECO:0000269|PubMed:16339187}. | |||||
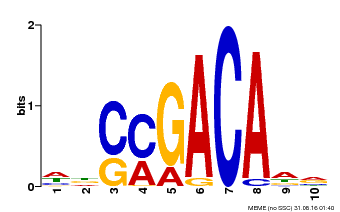
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024023134.1 | 0.0 | auxin response factor 2B isoform X1 | ||||
| Refseq | XP_024023135.1 | 0.0 | auxin response factor 2B isoform X2 | ||||
| Refseq | XP_024023136.1 | 0.0 | auxin response factor 2B isoform X3 | ||||
| Swissprot | Q94JM3 | 0.0 | ARFB_ARATH; Auxin response factor 2 | ||||
| TrEMBL | A0A2P5FI45 | 0.0 | A0A2P5FI45_TREOI; Auxin response factor | ||||
| STRING | XP_010099050.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3281 | 33 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||



