 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PK15677.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Cannabis
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 367aa MW: 41286.3 Da PI: 8.2878 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.4 | 2.8e-32 | 79 | 132 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtp+LH rFv+ave+LGG+e+AtPk +l+lm++kgL+++hvkSHLQ+YR+
PK15677.1 79 PRLRWTPDLHLRFVHAVERLGGQERATPKLVLQLMNIKGLSIAHVKSHLQMYRS 132
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 12.521 | 75 | 135 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.6E-30 | 75 | 133 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 9.14E-15 | 77 | 133 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 9.0E-23 | 79 | 134 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.4E-9 | 80 | 131 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 367 aa Download sequence Send to blast |
SISSSSSSGN DSEYSKTRLE YEEDEDEDHV YQSGEENKAN INGGLSSSNS TVEEEDNNDN 60 NKVNKSSSSV RPYVRSKMPR LRWTPDLHLR FVHAVERLGG QERATPKLVL QLMNIKGLSI 120 AHVKSHLQMY RSKKIDDAGQ VIGSHDQYHH HRHLVECGDR NIYNFSQLPM LQGYKQISHS 180 SSFRYGFGDP SWISTTSSFG NLRQYSRAGF YASSEAERKN ILAINNTNNA CNFIPSKNYN 240 ISSTISSIIN NHQHSSSWRN KAHNNKLIIK DIIDDHSSSL SPLIDQNSIA HLQVNKPKEL 300 TNFLTSSNNN VTHSDLVIST TTTSTSSMKR KASSDSNDLD LSLRLTTTTN TTHHNNAIVE 360 IHDHNKK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 7e-18 | 80 | 134 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 7e-18 | 80 | 134 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 9e-18 | 80 | 134 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 9e-18 | 80 | 134 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 9e-18 | 80 | 134 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 9e-18 | 80 | 134 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 7e-18 | 80 | 134 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 7e-18 | 80 | 134 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 7e-18 | 80 | 134 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 7e-18 | 80 | 134 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 7e-18 | 80 | 134 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 7e-18 | 80 | 134 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00297 | DAP | Transfer from AT2G38300 | Download |
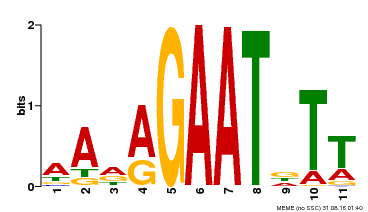
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010092813.1 | 6e-93 | two-component response regulator ORR22 | ||||
| TrEMBL | W9R2E6 | 1e-91 | W9R2E6_9ROSA; Putative Myb family transcription factor | ||||
| STRING | XP_010092813.1 | 2e-92 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2378 | 34 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38300.1 | 7e-54 | G2-like family protein | ||||



