 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PK12143.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Cannabis
|
||||||||
| Family | CPP | ||||||||
| Protein Properties | Length: 529aa MW: 57331.4 Da PI: 7.4856 | ||||||||
| Description | CPP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCR | 44.2 | 3.9e-14 | 42 | 81 | 2 | 42 |
TCR 2 ekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNkeek 42
++k+CnC++s+Clk+YCeCfa+g +C+ C+C +C N+ e+
PK12143.1 42 KQKQCNCRHSRCLKLYCECFASGIYCDG-CNCVNCYNNVEN 81
89**************************.********9875 PP
| |||||||
| 2 | TCR | 51.1 | 2.7e-16 | 126 | 165 | 1 | 40 |
TCR 1 kekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNke 40
k++kgC+Ckks ClkkYCeCf+a+ Cse+CkC dCkN e
PK12143.1 126 KHNKGCHCKKSGCLKKYCECFQANILCSENCKCMDCKNFE 165
589***********************************65 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01114 | 2.9E-16 | 41 | 81 | IPR033467 | Tesmin/TSO1-like CXC domain |
| PROSITE profile | PS51634 | 36.903 | 42 | 166 | IPR005172 | CRC domain |
| Pfam | PF03638 | 5.0E-11 | 43 | 78 | IPR005172 | CRC domain |
| SMART | SM01114 | 8.4E-20 | 126 | 167 | IPR033467 | Tesmin/TSO1-like CXC domain |
| Pfam | PF03638 | 9.9E-13 | 129 | 164 | IPR005172 | CRC domain |
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 529 aa Download sequence Send to blast |
XPPVVTGPVM PQPSVPSARI VKPDSPKSRS RPNEVKDGTP KKQKQCNCRH SRCLKLYCEC 60 FASGIYCDGC NCVNCYNNVE NEAARREAVE ATLERNPNAF RPKIASSPHG VRDNREDAGE 120 IMILGKHNKG CHCKKSGCLK KYCECFQANI LCSENCKCMD CKNFEGSEER QALFHGDHGS 180 NMAYIQQAAN AAITGAIGSS GYTSPPVSKK RKGQELFFGP TVTPKDTSVN RIGQFQQVNH 240 HRAPVPPSLP SVPAARLGSA TGVGPSKFTY RSLLADIIQP QDLKELCSVL VVVSGEAAKT 300 LAEQRSATEK RTEDQSETSL ASTTQDRLQN QKESDLEKSV DDCSSANQAD KTGADDSSSD 360 GADLPKGRPM SPDTLALMCD EQDTMFMAPA SPSGTGHGCN ISSQLPYEQG MPELYAEQER 420 LVLSRFRDCL KMVIALGEIK ETKYSSLAQS EPESHKDTLT NIGITNHKTE ISYHSGPAKN 480 GVLKTIVPPT TKTTQTISTA VAAVSNELPK IPQFPENGGI ANPKIEKEM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5fd3_A | 4e-32 | 34 | 175 | 2 | 132 | Protein lin-54 homolog |
| 5fd3_B | 4e-32 | 34 | 175 | 2 | 132 | Protein lin-54 homolog |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Plays a role in development of both male and female reproductive tissues. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00045 | PBM | Transfer from AT4G29000 | Download |
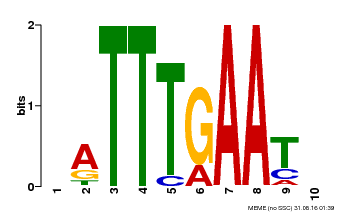
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015898233.1 | 0.0 | protein tesmin/TSO1-like CXC 5 | ||||
| Swissprot | Q9SZD1 | 0.0 | TCX5_ARATH; Protein tesmin/TSO1-like CXC 5 | ||||
| TrEMBL | A0A2P5FSX7 | 0.0 | A0A2P5FSX7_TREOI; Lin-54-like protein | ||||
| STRING | XP_010105053.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2626 | 33 | 79 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G29000.1 | 1e-165 | Tesmin/TSO1-like CXC domain-containing protein | ||||



