 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PK12073.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Cannabis
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 560aa MW: 61729.6 Da PI: 7.2948 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 52.5 | 8.6e-17 | 366 | 412 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn ErrRRdriN+++ L+el+P++ K +Ka++L +A+eY+ksLq
PK12073.1 366 VHNLSERRRRDRINEKMKALQELIPHS-----NKTDKASMLDEAIEYLKSLQ 412
6*************************7.....5******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 1.83E-20 | 359 | 426 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 9.9E-20 | 360 | 419 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 18.254 | 362 | 411 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 4.59E-18 | 365 | 416 | No hit | No description |
| Pfam | PF00010 | 3.5E-14 | 366 | 412 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 3.9E-18 | 368 | 417 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0009693 | Biological Process | ethylene biosynthetic process | ||||
| GO:0010244 | Biological Process | response to low fluence blue light stimulus by blue low-fluence system | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0010928 | Biological Process | regulation of auxin mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 560 aa Download sequence Send to blast |
MNNIPDWNFD CDLPVNGQKN PNGADHDLVE LLWRNGQVVL QSQTHRKASV GANESRQVQK 60 HDQQAIRNGG PYGNSMNMVQ DDDAVPWIHY PLEDAFEKDF CSNFFSELSS CDPIEIDKPI 120 RHVDEERLLS KFGGSDTTHV PSNSQHQNHQ KLKPSSNGVV PCPENHNMPP PRHNYSNSSV 180 HQKQNLGSLG KVVNFSQFSA MGKCDMRPQQ QQQAVKEPGN RGEVRECSAM TVGLSHCGSN 240 QLAAADLDVS WVSSNNGDGN NGLSAGAFKD DVQKILPRTR TSESGKAETL EPTLTSSSGG 300 SGSSFGRTCK QSTATTSANN NNNNKRKNRD AEELECQSDA AEHESAAANK SSQRSGSSRK 360 SRAAEVHNLS ERRRRDRINE KMKALQELIP HSNKTDKASM LDEAIEYLKS LQLQLQVMWM 420 GSGMAPMMFP GVQHYMSRLG MAMGPPSLPS IHNPMHLPRV PVVDQSMIGP QTTDQSVLCQ 480 TPAFNPINYQ NQMQNASFQE QYARYMGFHP MQTVSQPMNL FRFNPQTLQQ TQSIAQQQGI 540 STAPSNSGIP TNGALTGKMG |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 370 | 375 | ERRRRD |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00082 | ChIP-seq | Transfer from AT3G59060 | Download |
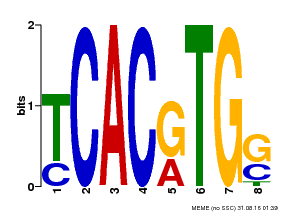
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010093193.2 | 0.0 | transcription factor PIF4 isoform X1 | ||||
| Refseq | XP_024019248.1 | 0.0 | transcription factor PIF4 isoform X2 | ||||
| TrEMBL | A0A2P5EVY1 | 0.0 | A0A2P5EVY1_TREOI; Basic helix-loop-helix transcription factor | ||||
| STRING | XP_010093193.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8146 | 29 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G59060.4 | 2e-36 | phytochrome interacting factor 3-like 6 | ||||



